Human paraneoplastic antigen Ma2 (PNMA2) forms icosahedral capsids that can be engineered for mRNA delivery
- PMID: 38437549
- PMCID: PMC10945824
- DOI: 10.1073/pnas.2307812120
Human paraneoplastic antigen Ma2 (PNMA2) forms icosahedral capsids that can be engineered for mRNA delivery
Abstract
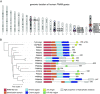
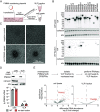
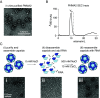
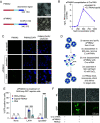
A number of endogenous genes in the human genome encode retroviral gag-like proteins, which were domesticated from ancient retroelements. The paraneoplastic Ma antigen (PNMA) family members encode a gag-like capsid domain, but their ability to assemble as capsids and traffic between cells remains mostly uncharacterized. Here, we systematically investigate human PNMA proteins and find that a number of PNMAs are secreted by human cells. We determine that PNMA2 forms icosahedral capsids efficiently but does not naturally encapsidate nucleic acids. We resolve the cryoelectron microscopy (cryo-EM) structure of PNMA2 and leverage the structure to design engineered PNMA2 (ePNMA2) particles with RNA packaging abilities. Recombinantly purified ePNMA2 proteins package mRNA molecules into icosahedral capsids and can function as delivery vehicles in mammalian cell lines, demonstrating the potential for engineered endogenous capsids as a nucleic acid therapy delivery modality.
Keywords: PNMA; VLP; capsid; gene therapy; mRNA delivery.
Conflict of interest statement
Competing interests statement:The authors declare no competing interest.
Figures

Similar articles
-
PNMA2 forms immunogenic non-enveloped virus-like capsids associated with paraneoplastic neurological syndrome.Cell. 2024 Feb 15;187(4):831-845.e19. doi: 10.1016/j.cell.2024.01.009. Epub 2024 Jan 31. Cell. 2024. PMID: 38301645 Free PMC article.
-
PNMA2 forms non-enveloped virus-like capsids that trigger paraneoplastic neurological syndrome.bioRxiv [Preprint]. 2023 Feb 9:2023.02.09.527862. doi: 10.1101/2023.02.09.527862. bioRxiv. 2023. Update in: Cell. 2024 Feb 15;187(4):831-845.e19. doi: 10.1016/j.cell.2024.01.009. PMID: 36798413 Free PMC article. Updated. Preprint.
-
PNMA2 mediates heterodimeric interactions and antagonizes chemo-sensitizing activities mediated by members of PNMA family.Biochem Biophys Res Commun. 2016 Apr 22;473(1):224-229. doi: 10.1016/j.bbrc.2016.03.083. Epub 2016 Mar 19. Biochem Biophys Res Commun. 2016. PMID: 27003254
-
Breaking Symmetry in Viral Icosahedral Capsids as Seen through the Lenses of X-ray Crystallography and Cryo-Electron Microscopy.Viruses. 2018 Feb 7;10(2):67. doi: 10.3390/v10020067. Viruses. 2018. PMID: 29414851 Free PMC article. Review.
-
Beyond structures of highly symmetric purified viral capsids by cryo-EM.Curr Opin Struct Biol. 2018 Oct;52:25-31. doi: 10.1016/j.sbi.2018.07.011. Epub 2018 Aug 7. Curr Opin Struct Biol. 2018. PMID: 30096461 Review.
Cited by
-
The retrotransposon-derived capsid genes PNMA1 and PNMA4 maintain reproductive capacity.Res Sq [Preprint]. 2024 Jul 10:rs.3.rs-4559920. doi: 10.21203/rs.3.rs-4559920/v1. Res Sq. 2024. PMID: 39041030 Free PMC article. Preprint.
-
The myth of high-resolution liquid phase biological electron microscopy.Protein Sci. 2024 Aug;33(8):e5125. doi: 10.1002/pro.5125. Protein Sci. 2024. PMID: 39037286 Free PMC article.
-
Biomimetic Nucleic Acid Drug Delivery Systems for Relieving Tumor Immunosuppressive Microenvironment.Pharmaceutics. 2024 Aug 1;16(8):1028. doi: 10.3390/pharmaceutics16081028. Pharmaceutics. 2024. PMID: 39204373 Free PMC article. Review.
-
RNA delivery systems.Proc Natl Acad Sci U S A. 2024 Mar 12;121(11):e2315789121. doi: 10.1073/pnas.2315789121. Epub 2024 Mar 4. Proc Natl Acad Sci U S A. 2024. PMID: 38437565 Free PMC article. No abstract available.
-
Evolution of Virus-like Features and Intrinsically Disordered Regions in Retrotransposon-derived Mammalian Genes.Mol Biol Evol. 2024 Aug 2;41(8):msae154. doi: 10.1093/molbev/msae154. Mol Biol Evol. 2024. PMID: 39101471 Free PMC article.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

