Microbiome and plant cell transformation trigger insect gall induction in cassava
- PMID: 38126017
- PMCID: PMC10731979
- DOI: 10.3389/fpls.2023.1237966
Microbiome and plant cell transformation trigger insect gall induction in cassava
Abstract
Several specialised insects can manipulate normal plant development to induce a highly organised structure known as a gall, which represents one of the most complex interactions between insects and plants. Thus far, the mechanism for insect-induced plant galls has remained elusive. To study the induction mechanism of insect galls, we selected the gall induced by Iatrophobia brasiliensis (Diptera: Cecidomyiidae) in cassava (Euphorbiaceae: Manihot esculenta Crantz) as our model. PCR-based molecular markers and deep metagenomic sequencing data were employed to analyse the gall microbiome and to test the hypothesis that gall cells are genetically transformed by insect vectored bacteria. A shotgun sequencing discrimination approach was implemented to selectively discriminate between foreign DNA and the reference host plant genome. Several known candidate insertion sequences were identified, the most significant being DNA sequences found in bacterial genes related to the transcription regulatory factor CadR, cadmium-transporting ATPase encoded by the cadA gene, nitrate transport permease protein (nrtB gene), and arsenical pump ATPase (arsA gene). In addition, a DNA fragment associated with ubiquitin-like gene E2 was identified as a potential accessory genetic element involved in gall induction mechanism. Furthermore, our results suggest that the increased quality and rapid development of gall tissue are mostly driven by microbiome enrichment and the acquisition of critical endophytes. An initial gall-like structure was experimentally obtained in M. esculenta cultured tissues through inoculation assays using a Rhodococcus bacterial strain that originated from the inducing insect, which we related to the gall induction process. We provide evidence that the modification of the endophytic microbiome and the genetic transformation of plant cells in M. esculenta are two essential requirements for insect-induced gall formation. Based on these findings and having observed the same potential DNA marker in galls from other plant species (ubiquitin-like gene E2), we speculate that bacterially mediated genetic transformation of plant cells may represent a more widespread gall induction mechanism found in nature.
Keywords: Iatrophobia brasiliensis; Manihot esculenta; endophytes; genetic transformation; induction mechanism; metagenomics; plant galls.
Copyright © 2023 Gätjens-Boniche, Jiménez-Madrigal, Whetten, Valenzuela-Diaz, Alemán-Gutiérrez, Hanson and Pinto-Tomás.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures




Similar articles
-
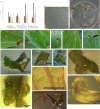
Nutritive tissue rich in reserves in the cell wall and protoplast: the case of Manihot esculenta (Euphorbiaceae) galls induced by Iatrophobia brasiliensis (Diptera, Cecidomyiidae).Protoplasma. 2024 May;261(3):513-525. doi: 10.1007/s00709-023-01912-z. Epub 2023 Dec 19. Protoplasma. 2024. PMID: 38114665
-
Manipulation of host plant cells and tissues by gall-inducing insects and adaptive strategies used by different feeding guilds.J Insect Physiol. 2016 Jan;84:103-113. doi: 10.1016/j.jinsphys.2015.11.012. Epub 2015 Nov 24. J Insect Physiol. 2016. PMID: 26620152 Review.
-
Comparative transcriptome analysis of galls from four different host plants suggests the molecular mechanism of gall development.PLoS One. 2019 Oct 24;14(10):e0223686. doi: 10.1371/journal.pone.0223686. eCollection 2019. PLoS One. 2019. PMID: 31647845 Free PMC article.
-
Recent Progress Regarding the Molecular Aspects of Insect Gall Formation.Int J Mol Sci. 2021 Aug 30;22(17):9424. doi: 10.3390/ijms22179424. Int J Mol Sci. 2021. PMID: 34502330 Free PMC article. Review.
-
Cytokinins Are Abundant and Widespread Among Insect Species.Plants (Basel). 2020 Feb 6;9(2):208. doi: 10.3390/plants9020208. Plants (Basel). 2020. PMID: 32041320 Free PMC article.
References
-
- Alexa A. (2022). Rahnenfuhrer J. topGO: Enrichment analysis for gene ontology (2.46. 0) [Computer software]. Bioconductor version: release (3.14).
-
- Ananthakrishnan T. (1998). Insect gall systems: patterns, processes and adaptive diversity. Curr. Sci. 75 (7), 672–676.
Grants and funding
LinkOut - more resources
Full Text Sources

