Studies of Genetic and Proteomic Risk Factors of Amyotrophic Lateral Sclerosis Inspire Biomarker Development and Gene Therapy
- PMID: 37566027
- PMCID: PMC10417729
- DOI: 10.3390/cells12151948
Studies of Genetic and Proteomic Risk Factors of Amyotrophic Lateral Sclerosis Inspire Biomarker Development and Gene Therapy
Abstract
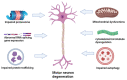
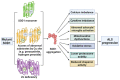
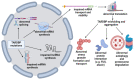
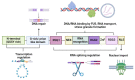
Amyotrophic lateral sclerosis (ALS) is an incurable neurodegenerative disease affecting the upper and lower motor neurons, leading to muscle weakness, motor impairments, disabilities and death. Approximately 5-10% of ALS cases are associated with positive family history (familial ALS or fALS), whilst the remainder are sporadic (sporadic ALS, sALS). At least 50 genes have been identified as causative or risk factors for ALS. Established pathogenic variants include superoxide dismutase type 1 (SOD1), chromosome 9 open reading frame 72 (c9orf72), TAR DNA Binding Protein (TARDBP), and Fused In Sarcoma (FUS); additional ALS-related genes including Charged Multivesicular Body Protein 2B (CHMP2B), Senataxin (SETX), Sequestosome 1 (SQSTM1), TANK Binding Kinase 1 (TBK1) and NIMA Related Kinase 1 (NEK1), have been identified. Mutations in these genes could impair different mechanisms, including vesicle transport, autophagy, and cytoskeletal or mitochondrial functions. So far, there is no effective therapy against ALS. Thus, early diagnosis and disease risk predictions remain one of the best options against ALS symptomologies. Proteomic biomarkers, microRNAs, and extracellular vehicles (EVs) serve as promising tools for disease diagnosis or progression assessment. These markers are relatively easy to obtain from blood or cerebrospinal fluids and can be used to identify potential genetic causative and risk factors even in the preclinical stage before symptoms appear. In addition, antisense oligonucleotides and RNA gene therapies have successfully been employed against other diseases, such as childhood-onset spinal muscular atrophy (SMA), which could also give hope to ALS patients. Therefore, an effective gene and biomarker panel should be generated for potentially "at risk" individuals to provide timely interventions and better treatment outcomes for ALS patients as soon as possible.
Keywords: amyotrophic lateral sclerosis; biomarkers; diagnosis; genetic factors; proteomic markers; therapy.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
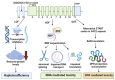

Similar articles
-
Genetics screening in an Italian cohort of patients with Amyotrophic Lateral Sclerosis: the importance of early testing and its implication.J Neurol. 2024 Apr;271(4):1921-1936. doi: 10.1007/s00415-023-12142-x. Epub 2023 Dec 19. J Neurol. 2024. PMID: 38112783
-
Amyotrophic Lateral Sclerosis Overview.2001 Mar 23 [updated 2023 Sep 28]. In: Adam MP, Feldman J, Mirzaa GM, Pagon RA, Wallace SE, Amemiya A, editors. GeneReviews® [Internet]. Seattle (WA): University of Washington, Seattle; 1993–2025. 2001 Mar 23 [updated 2023 Sep 28]. In: Adam MP, Feldman J, Mirzaa GM, Pagon RA, Wallace SE, Amemiya A, editors. GeneReviews® [Internet]. Seattle (WA): University of Washington, Seattle; 1993–2025. PMID: 20301623 Free Books & Documents. Review.
-
Accuracy of a machine learning method based on structural and locational information from AlphaFold2 for predicting the pathogenicity of TARDBP and FUS gene variants in ALS.BMC Bioinformatics. 2023 May 19;24(1):206. doi: 10.1186/s12859-023-05338-5. BMC Bioinformatics. 2023. PMID: 37208601 Free PMC article.
-
Phenotype difference between ALS patients with expanded repeats in C9ORF72 and patients with mutations in other ALS-related genes.J Med Genet. 2012 Apr;49(4):258-63. doi: 10.1136/jmedgenet-2011-100699. J Med Genet. 2012. PMID: 22499346
-
Pathogenic commonalities between spinal muscular atrophy and amyotrophic lateral sclerosis: Converging roads to therapeutic development.Eur J Med Genet. 2018 Nov;61(11):685-698. doi: 10.1016/j.ejmg.2017.12.001. Epub 2017 Dec 5. Eur J Med Genet. 2018. PMID: 29313812 Review.
Cited by
-
CNS cell-derived exosome signatures as blood-based biomarkers of neurodegenerative diseases.Front Neurosci. 2024 Jun 20;18:1426700. doi: 10.3389/fnins.2024.1426700. eCollection 2024. Front Neurosci. 2024. PMID: 38966760 Free PMC article. Review.
-
The role of ALDH2 rs671 polymorphism and C-reactive protein in the phenotypes of male ALS patients.Front Neurosci. 2024 Sep 3;18:1397991. doi: 10.3389/fnins.2024.1397991. eCollection 2024. Front Neurosci. 2024. PMID: 39290715 Free PMC article.
References
-
- Hulisz D. Amyotrophic lateral sclerosis: Disease state overview. Am. J. Manag. Care. 2018;24((Suppl. S15)):S320–S326. - PubMed
Publication types
MeSH terms
Substances
Supplementary concepts
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

