Copper metabolism in cell death and autophagy
- PMID: 37055935
- PMCID: PMC10351475
- DOI: 10.1080/15548627.2023.2200554
Copper metabolism in cell death and autophagy
Abstract
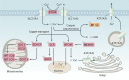
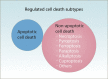
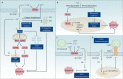
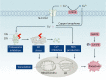
Copper is an essential trace element in biological systems, maintaining the activity of enzymes and the function of transcription factors. However, at high concentrations, copper ions show increased toxicity by inducing regulated cell death, such as apoptosis, paraptosis, pyroptosis, ferroptosis, and cuproptosis. Furthermore, copper ions can trigger macroautophagy/autophagy, a lysosome-dependent degradation pathway that plays a dual role in regulating the survival or death fate of cells under various stress conditions. Pathologically, impaired copper metabolism due to environmental or genetic causes is implicated in a variety of human diseases, such as rare Wilson disease and common cancers. Therapeutically, copper-based compounds are potential chemotherapeutic agents that can be used alone or in combination with other drugs or approaches to treat cancer. Here, we review the progress made in understanding copper metabolic processes and their impact on the regulation of cell death and autophagy. This knowledge may help in the design of future clinical tools to improve cancer diagnosis and treatment.Abbreviations: ACSL4, acyl-CoA synthetase long chain family member 4; AIFM1/AIF, apoptosis inducing factor mitochondria associated 1; AIFM2, apoptosis inducing factor mitochondria associated 2; ALDH, aldehyde dehydrogenase; ALOX, arachidonate lipoxygenase; AMPK, AMP-activated protein kinase; APAF1, apoptotic peptidase activating factor 1; ATF4, activating transcription factor 4; ATG, autophagy related; ATG13, autophagy related 13; ATG5, autophagy related 5; ATOX1, antioxidant 1 copper chaperone; ATP, adenosine triphosphate; ATP7A, ATPase copper transporting alpha; ATP7B, ATPase copper transporting beta; BAK1, BCL2 antagonist/killer 1; BAX, BCL2 associated X apoptosis regulator; BBC3/PUMA, BCL2 binding component 3; BCS, bathocuproinedisulfonic acid; BECN1, beclin 1; BID, BH3 interacting domain death agonist; BRCA1, BRCA1 DNA repair associated; BSO, buthionine sulphoximine; CASP1, caspase 1; CASP3, caspase 3; CASP4/CASP11, caspase 4; CASP5, caspase 5; CASP8, caspase 8; CASP9, caspase 9; CCS, copper chaperone for superoxide dismutase; CD274/PD-L1, CD274 molecule; CDH2, cadherin 2; CDKN1A/p21, cyclin dependent kinase inhibitor 1A; CDKN1B/p27, cyclin-dependent kinase inhibitor 1B; COMMD10, COMM domain containing 10; CoQ10, coenzyme Q 10; CoQ10H2, reduced coenzyme Q 10; COX11, cytochrome c oxidase copper chaperone COX11; COX17, cytochrome c oxidase copper chaperone COX17; CP, ceruloplasmin; CYCS, cytochrome c, somatic; DBH, dopamine beta-hydroxylase; DDIT3/CHOP, DNA damage inducible transcript 3; DLAT, dihydrolipoamide S-acetyltransferase; DTC, diethyldithiocarbamate; EIF2A, eukaryotic translation initiation factor 2A; EIF2AK3/PERK, eukaryotic translation initiation factor 2 alpha kinase 3; ER, endoplasmic reticulum; ESCRT-III, endosomal sorting complex required for transport-III; ETC, electron transport chain; FABP3, fatty acid binding protein 3; FABP7, fatty acid binding protein 7; FADD, Fas associated via death domain; FAS, Fas cell surface death receptor; FASL, Fas ligand; FDX1, ferredoxin 1; GNAQ/11, G protein subunit alpha q/11; GPX4, glutathione peroxidase 4; GSDMD, gasdermin D; GSH, glutathione; HDAC, histone deacetylase; HIF1, hypoxia inducible factor 1; HIF1A, hypoxia inducible factor 1 subunit alpha; HMGB1, high mobility group box 1; IL1B, interleukin 1 beta; IL17, interleukin 17; KRAS, KRAS proto-oncogene, GTPase; LOX, lysyl oxidase; LPCAT3, lysophosphatidylcholine acyltransferase 3; MAP1LC3, microtubule associated protein 1 light chain 3; MAP2K1, mitogen-activated protein kinase kinase 1; MAP2K2, mitogen-activated protein kinase kinase 2; MAPK, mitogen-activated protein kinases; MAPK14/p38, mitogen-activated protein kinase 14; MEMO1, mediator of cell motility 1; MT-CO1/COX1, mitochondrially encoded cytochrome c oxidase I; MT-CO2/COX2, mitochondrially encoded cytochrome c oxidase II; MTOR, mechanistic target of rapamycin kinase; MTs, metallothioneins; NAC, N-acetylcysteine; NFKB/NF-Κb, nuclear factor kappa B; NLRP3, NLR family pyrin domain containing 3; NPLOC4/NPL4, NPL4 homolog ubiquitin recognition factor; PDE3B, phosphodiesterase 3B; PDK1, phosphoinositide dependent protein kinase 1; PHD, prolyl-4-hydroxylase domain; PIK3C3/VPS34, phosphatidylinositol 3-kinase catalytic subunit type 3; PMAIP1/NOXA, phorbol-12-myristate-13-acetate-induced protein 1; POR, cytochrome P450 oxidoreductase; PUFA-PL, PUFA of phospholipids; PUFAs, polyunsaturated fatty acids; ROS, reactive oxygen species; SCO1, synthesis of cytochrome C oxidase 1; SCO2, synthesis of cytochrome C oxidase 2; SLC7A11, solute carrier family 7 member 11; SLC11A2/DMT1, solute carrier family 11 member 2; SLC31A1/CTR1, solute carrier family 31 member 1; SLC47A1, solute carrier family 47 member 1; SOD1, superoxide dismutase; SP1, Sp1 transcription factor; SQSTM1/p62, sequestosome 1; STEAP4, STEAP4 metalloreductase; TAX1BP1, Tax1 binding protein 1; TEPA, tetraethylenepentamine; TFEB, transcription factor EB; TM, tetrathiomolybdate; TP53/p53, tumor protein p53; TXNRD1, thioredoxin reductase 1; UCHL5, ubiquitin C-terminal hydrolase L5; ULK1, Unc-51 like autophagy activating kinase 1; ULK1, unc-51 like autophagy activating kinase 1; ULK2, unc-51 like autophagy activating kinase 2; USP14, ubiquitin specific peptidase 14; VEGF, vascular endothelial gro wth factor; XIAP, X-linked inhibitor of apoptosis.
Keywords: autophagy; cancer; cell death; copper; cuproptosis; ferroptosis.
Conflict of interest statement
No potential conflict of interest was reported by the authors.
Figures

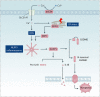
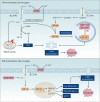
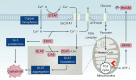
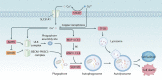
Similar articles
-
Autophagy in the physiological endometrium and cancer.Autophagy. 2021 May;17(5):1077-1095. doi: 10.1080/15548627.2020.1752548. Epub 2020 May 13. Autophagy. 2021. PMID: 32401642 Free PMC article. Review.
-
How autophagy controls the intestinal epithelial barrier.Autophagy. 2022 Jan;18(1):86-103. doi: 10.1080/15548627.2021.1909406. Epub 2021 Apr 27. Autophagy. 2022. PMID: 33906557 Free PMC article. Review.
-
A systematic review of p53 regulation of oxidative stress in skeletal muscle.Redox Rep. 2018 Dec;23(1):100-117. doi: 10.1080/13510002.2017.1416773. Epub 2018 Jan 3. Redox Rep. 2018. PMID: 29298131 Free PMC article. Review.
-
Organelle-specific autophagy in inflammatory diseases: a potential therapeutic target underlying the quality control of multiple organelles.Autophagy. 2021 Feb;17(2):385-401. doi: 10.1080/15548627.2020.1725377. Epub 2020 Feb 12. Autophagy. 2021. PMID: 32048886 Free PMC article. Review.
-
Impaired autophagy and APP processing in Alzheimer's disease: The potential role of Beclin 1 interactome.Prog Neurobiol. 2013 Jul-Aug;106-107:33-54. doi: 10.1016/j.pneurobio.2013.06.002. Epub 2013 Jul 1. Prog Neurobiol. 2013. PMID: 23827971 Review.
Cited by
-
The Role of Epigenetic Control of Mitochondrial (Dys)Function in MASLD Onset and Progression.Nutrients. 2023 Nov 12;15(22):4757. doi: 10.3390/nu15224757. Nutrients. 2023. PMID: 38004151 Free PMC article. Review.
-
An omics review and perspective of researches on intrahepatic cholestasis of pregnancy.Front Endocrinol (Lausanne). 2024 Jan 8;14:1267195. doi: 10.3389/fendo.2023.1267195. eCollection 2023. Front Endocrinol (Lausanne). 2024. PMID: 38260124 Free PMC article. Review.
-
miR-137: a potential therapeutic target for lung cancer.Front Cell Dev Biol. 2024 Aug 23;12:1427724. doi: 10.3389/fcell.2024.1427724. eCollection 2024. Front Cell Dev Biol. 2024. PMID: 39247624 Free PMC article. Review.
-
Iron and copper: critical executioners of ferroptosis, cuproptosis and other forms of cell death.Cell Commun Signal. 2023 Nov 16;21(1):327. doi: 10.1186/s12964-023-01267-1. Cell Commun Signal. 2023. PMID: 37974196 Free PMC article. Review.
-
Research progress on ferroptosis in the pathogenesis and treatment of neurodegenerative diseases.Front Cell Neurosci. 2024 Mar 7;18:1359453. doi: 10.3389/fncel.2024.1359453. eCollection 2024. Front Cell Neurosci. 2024. PMID: 38515787 Free PMC article. Review.
References
-
- Chen J, Jiang Y, Shi H, et al. The molecular mechanisms of copper metabolism and its roles in human diseases. Pflügers Archiv - European Journal of Physiology. 2020;472(10):1415–1429. - PubMed
-
- Steffens GC, Biewald R, Buse G.. Cytochrome c oxidase is a three-copper, two-heme-A protein. Eur J Biochem. 1987;164(2):295–300. - PubMed
-
- In: Harris ED. Copper as acofactor and regulator of copper,zinc superoxide dismutase. J Nutr 1992. Vol. 122pp. 636–640 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous
