The molecular biology of tubulinopathies: Understanding the impact of variants on tubulin structure and microtubule regulation
- PMID: 36406756
- PMCID: PMC9666403
- DOI: 10.3389/fncel.2022.1023267
The molecular biology of tubulinopathies: Understanding the impact of variants on tubulin structure and microtubule regulation
Abstract
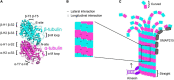
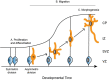
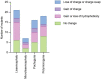
Heterozygous, missense mutations in both α- and β-tubulin genes have been linked to an array of neurodevelopment disorders, commonly referred to as "tubulinopathies." To date, tubulinopathy mutations have been identified in three β-tubulin isotypes and one α-tubulin isotype. These mutations occur throughout the different genetic domains and protein structures of these tubulin isotypes, and the field is working to address how this molecular-level diversity results in different cellular and tissue-level pathologies. Studies from many groups have focused on elucidating the consequences of individual mutations; however, the field lacks comprehensive models for the molecular etiology of different types of tubulinopathies, presenting a major gap in diagnosis and treatment. This review highlights recent advances in understanding tubulin structural dynamics, the roles microtubule-associated proteins (MAPs) play in microtubule regulation, and how these are inextricably linked. We emphasize the value of investigating interactions between tubulin structures, microtubules, and MAPs to understand and predict the impact of tubulinopathy mutations at the cell and tissue levels. Microtubule regulation is multifaceted and provides a complex set of controls for generating a functional cytoskeleton at the right place and right time during neurodevelopment. Understanding how tubulinopathy mutations disrupt distinct subsets of those controls, and how that ultimately disrupts neurodevelopment, will be important for establishing mechanistic themes among tubulinopathies that may lead to insights in other neurodevelopment disorders and normal neurodevelopment.
Keywords: cytoskeleton; microtubule; neurodevelopment; protein structure; tubulinopathy.
Copyright © 2022 Hoff, Neumann and Moore.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures



Similar articles
-
TUBA1A tubulinopathy mutants disrupt neuron morphogenesis and override XMAP215/Stu2 regulation of microtubule dynamics.Elife. 2022 May 5;11:e76189. doi: 10.7554/eLife.76189. Elife. 2022. PMID: 35511030 Free PMC article.
-
Tubulin mutations in brain development disorders: Why haploinsufficiency does not explain TUBA1A tubulinopathies.Cytoskeleton (Hoboken). 2020 Mar;77(3-4):40-54. doi: 10.1002/cm.21567. Epub 2019 Oct 31. Cytoskeleton (Hoboken). 2020. PMID: 31574570
-
Insights on the Role of α- and β-Tubulin Isotypes in Early Brain Development.Mol Neurobiol. 2023 Jul;60(7):3803-3823. doi: 10.1007/s12035-023-03302-1. Epub 2023 Mar 21. Mol Neurobiol. 2023. PMID: 36943622 Review.
-
Kinetically Stabilizing Mutations in Beta Tubulins Create Isotype-Specific Brain Malformations.Front Cell Dev Biol. 2021 Nov 18;9:765992. doi: 10.3389/fcell.2021.765992. eCollection 2021. Front Cell Dev Biol. 2021. PMID: 34869359 Free PMC article.
-
Tubulin isotypes - functional insights from model organisms.J Cell Sci. 2022 May 1;135(9):jcs259539. doi: 10.1242/jcs.259539. Epub 2022 May 6. J Cell Sci. 2022. PMID: 35522156 Review.
Cited by
-
Interdisciplinary fetal-neonatal neurology training applies neural exposome perspectives to neurology principles and practice.Front Neurol. 2024 Jan 15;14:1321674. doi: 10.3389/fneur.2023.1321674. eCollection 2023. Front Neurol. 2024. PMID: 38288328 Free PMC article. Review.
-
Taxol acts differently on different tubulin isotypes.Commun Biol. 2023 Sep 16;6(1):946. doi: 10.1038/s42003-023-05306-y. Commun Biol. 2023. PMID: 37717119 Free PMC article.
-
Microtubule-associated protein MAP7 promotes tubulin posttranslational modifications and cargo transport to enable osmotic adaptation.Dev Cell. 2024 Jun 17;59(12):1553-1570.e7. doi: 10.1016/j.devcel.2024.03.022. Epub 2024 Apr 3. Dev Cell. 2024. PMID: 38574732
-
Tubulin CFEOM mutations both inhibit or activate kinesin motor activity.Mol Biol Cell. 2024 Mar 1;35(3):ar32. doi: 10.1091/mbc.E23-01-0020. Epub 2024 Jan 3. Mol Biol Cell. 2024. PMID: 38170592 Free PMC article.
-
A Variant in TBCD Associated with Motoneuronopathy and Corpus Callosum Hypoplasia: A Case Report.Int J Mol Sci. 2023 Aug 3;24(15):12386. doi: 10.3390/ijms241512386. Int J Mol Sci. 2023. PMID: 37569761 Free PMC article.
References
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

