Dynamic Diatom-Bacteria Consortia in Synthetic Plankton Communities
- PMID: 36300970
- PMCID: PMC9680611
- DOI: 10.1128/aem.01619-22
Dynamic Diatom-Bacteria Consortia in Synthetic Plankton Communities
Abstract
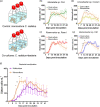
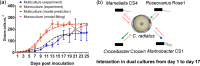
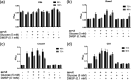
Microalgae that form phytoplankton live and die in a complex microbial consortium in which they co-exist with bacteria and other microorganisms. The dynamics of species succession in the plankton depends on the interplay of these partners. Bacteria utilize substrates produced by the phototrophic algae, while algal growth can be supported by bacterial exudates. Bacteria might also use chemical mediators with algicidal properties to attack algae. To elucidate whether specific bacteria play universal or context-specific roles in the interaction with phytoplankton, we investigated the effect of cocultured bacteria on the growth of 8 microalgae. An interaction matrix revealed that the function of a given bacterium is highly dependent on the cocultured partner. We observed no universally algicidal or universally growth-promoting bacteria. The activity of bacteria can even change during the aging of an algal culture from inhibitory to stimulatory or vice versa. We further established a synthetic phytoplankton/bacteria community with the centric diatom, Coscinodiscus radiatus, and 4 phylogenetically distinctive bacterial isolates, Mameliella sp., Roseovarius sp., Croceibacter sp., and Marinobacter sp. Supported by a Lotka-Volterra model, we show that interactions within the consortium are specific and that the sum of the pairwise interactions can explain algal and bacterial growth in the community. No synergistic effects between bacteria in the presence of the diatom was observed. Our survey documents highly species-specific interactions that are dependent on algal fitness, bacterial metabolism, and community composition. This species specificity may underly the high complexity of the multi-species plankton communities observed in nature. IMPORTANCE The marine food web is fueled by phototrophic phytoplankton. These algae are central primary producers responsible for the fixation of ca. 40% of the global CO2. Phytoplankton always co-occur with a diverse bacterial community in nature. This diversity suggests the existence of ecological niches for the associated bacteria. We show that the interaction between algae and bacteria is highly species-specific. Furthermore, both, the fitness stage of the algae and the community composition are relevant in determining the effect of bacteria on algal growth. We conclude that bacteria should not be sorted into algicidal or growth supporting categories; instead, a context-specific function of the bacteria in the plankton must be considered. This functional diversity of single players within a consortium may underly the observed diversity in the plankton.
Keywords: bacteria; consortia; diatoms; host-cell interactions; interactions; microalgae; microbiome; phytoplankton.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


Similar articles
-
The Algicidal Bacterium Kordia algicida Shapes a Natural Plankton Community.Appl Environ Microbiol. 2019 Mar 22;85(7):e02779-18. doi: 10.1128/AEM.02779-18. Print 2019 Apr 1. Appl Environ Microbiol. 2019. PMID: 30737345 Free PMC article.
-
Algicidal bacteria trigger contrasting responses in model diatom communities of different composition.Microbiologyopen. 2019 Aug;8(8):e00818. doi: 10.1002/mbo3.818. Epub 2019 Feb 27. Microbiologyopen. 2019. PMID: 30809963 Free PMC article.
-
Algal Oxylipins Mediate the Resistance of Diatoms against Algicidal Bacteria.Mar Drugs. 2018 Dec 4;16(12):486. doi: 10.3390/md16120486. Mar Drugs. 2018. PMID: 30518148 Free PMC article.
-
Strategies and ecological roles of algicidal bacteria.FEMS Microbiol Rev. 2017 Nov 1;41(6):880-899. doi: 10.1093/femsre/fux029. FEMS Microbiol Rev. 2017. PMID: 28961821 Review.
-
Temporal and Spatial Signaling Mediating the Balance of the Plankton Microbiome.Ann Rev Mar Sci. 2022 Jan 3;14:239-260. doi: 10.1146/annurev-marine-042021-012353. Epub 2021 Aug 26. Ann Rev Mar Sci. 2022. PMID: 34437810 Review.
Cited by
-
Abundant Sulfitobacter marine bacteria protect Emiliania huxleyi algae from pathogenic bacteria.ISME Commun. 2023 Sep 22;3(1):100. doi: 10.1038/s43705-023-00311-y. ISME Commun. 2023. PMID: 37740057 Free PMC article.
-
Temporal variability in the growth-enhancing effects of different bacteria within the microbiome of the diatom Actinocyclus sp.Front Microbiol. 2023 Aug 7;14:1230349. doi: 10.3389/fmicb.2023.1230349. eCollection 2023. Front Microbiol. 2023. PMID: 37608955 Free PMC article.
-
Forensic Diatom Analysis: Where Do We Stand and What Are the Latest Diagnostic Advances?Diagnostics (Basel). 2024 Oct 16;14(20):2302. doi: 10.3390/diagnostics14202302. Diagnostics (Basel). 2024. PMID: 39451625 Free PMC article. Review.
-
Heterotrophic bacteria trigger transcriptome remodelling in the photosynthetic picoeukaryote Micromonas commoda.Environ Microbiol Rep. 2024 Jun;16(3):e13285. doi: 10.1111/1758-2229.13285. Environ Microbiol Rep. 2024. PMID: 38778545 Free PMC article.
-
Bacteria modulate microalgal aging physiology through the induction of extracellular vesicle production to remove harmful metabolites.Nat Microbiol. 2024 Sep;9(9):2356-2368. doi: 10.1038/s41564-024-01746-2. Epub 2024 Aug 14. Nat Microbiol. 2024. PMID: 39143356 Free PMC article.
References
-
- Khangaonkar T, Sackmann B, Long W, Mohamedali T, Roberts M. 2012. Simulation of annual biogeochemical cycles of nutrient balance, phytoplankton bloom(s), and DO in Puget Sound using an unstructured grid model. Ocean Dynamics 62:1353–1379. 10.1007/s10236-012-0562-4. - DOI
-
- Litchman E, Klausmeier CA. 2008. Trait-based community ecology of phytoplankton. Annu Rev Ecol Evol Syst 39:615–639. 10.1146/annurev.ecolsys.39.110707.173549. - DOI
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources

