Genomic Analysis of an Excellent Wine-Making Strain Oenococcus oeni SD-2a
- PMID: 35716166
- PMCID: PMC9252139
- DOI: 10.33073/pjm-2022-026
Genomic Analysis of an Excellent Wine-Making Strain Oenococcus oeni SD-2a
Abstract
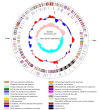
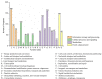
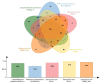
Oenococcus oeni is an important microorganism in wine-making-related engineering, and it improves wine quality and stability through malolactic fermentation. Although the genomes of more than 200 O. oeni strains have been sequenced, only a few include completed genome maps. Here, the genome sequence of O. oeni SD-2a, isolated from Shandong, China, has been determined. It is a fully assembled genome sequence of this strain. The complete genome is 1,989,703 bp with a G+C content of 37.8% without a plasmid. The genome includes almost all the essential genes involved in central metabolic pathways and the stress genes reported in other O. oeni strains. Some natural competence-related genes, like comEA, comEC, comFA, comG operon, and comFC, suggest that O. oeni SD-2a may have natural transformation potential. A comparative genomics analysis revealed 730 gene clusters in O. oeni SD-2a homologous to those in four other lactic acid bacteria species (O. oeni PSU-1, O. oeni CRBO-11381, Lactiplantibacillus plantarum UNQLp11, and Pediococcus pentosaceus KCCM40703). A collinearity analysis showed poor collinearity between O. oeni SD-2a and O. oeni PSU-1, indicating great differences in their evolutionary histories. The results provide general knowledge of O. oeni SD-2a and lay the foundation for specific gene function analyses.
Keywords: Oenococcus oeni SD-2a; collinearity; genome; homologous genes.
© 2022 Longxiang Liu et al., published by Sciendo.
Conflict of interest statement
The authors do not report any financial or personal connections with other persons or organizations, which might negatively affect the contents of this publication and/or claim authorship rights to this publication.
Figures


Similar articles
-
Identification of variable genomic regions related to stress response in Oenococcus oeni.Food Res Int. 2017 Dec;102:625-638. doi: 10.1016/j.foodres.2017.09.039. Epub 2017 Sep 19. Food Res Int. 2017. PMID: 29195994
-
Expanding the biodiversity of Oenococcus oeni through comparative genomics of apple cider and kombucha strains.BMC Genomics. 2019 May 2;20(1):330. doi: 10.1186/s12864-019-5692-3. BMC Genomics. 2019. PMID: 31046679 Free PMC article.
-
Genomic analysis of Oenococcus oeni PSU-1 and its relevance to winemaking.FEMS Microbiol Rev. 2005 Aug;29(3):465-75. doi: 10.1016/j.femsre.2005.04.011. FEMS Microbiol Rev. 2005. PMID: 16125008 Review.
-
Distribution of Oenococcus oeni populations in natural habitats.Appl Microbiol Biotechnol. 2019 Apr;103(7):2937-2945. doi: 10.1007/s00253-019-09689-z. Epub 2019 Feb 20. Appl Microbiol Biotechnol. 2019. PMID: 30788540 Free PMC article. Review.
-
Comparative functional analysis of malate metabolism genes in Oenococcus oeni and Lactiplantibacillus plantarum at low pH and their roles in acid stress response.Food Res Int. 2022 Jul;157:111235. doi: 10.1016/j.foodres.2022.111235. Epub 2022 Apr 14. Food Res Int. 2022. PMID: 35761547
Cited by
-
Heterologous expression of the Oenococcus oeni two-component signal transduction response regulator in the Lactiplantibacillus plantarum WCFS1 strain enhances acid stress tolerance.BMC Microbiol. 2024 Sep 28;24(1):370. doi: 10.1186/s12866-024-03498-9. BMC Microbiol. 2024. PMID: 39342090 Free PMC article.
References
MeSH terms
Supplementary concepts
Grants and funding
LinkOut - more resources
Full Text Sources
