Open Issues for Protein Function Assignment in Haloferax volcanii and Other Halophilic Archaea
- PMID: 34202810
- PMCID: PMC8305020
- DOI: 10.3390/genes12070963
Open Issues for Protein Function Assignment in Haloferax volcanii and Other Halophilic Archaea
Abstract
Background: Annotation ambiguities and annotation errors are a general challenge in genomics. While a reliable protein function assignment can be obtained by experimental characterization, this is expensive and time-consuming, and the number of such Gold Standard Proteins (GSP) with experimental support remains very low compared to proteins annotated by sequence homology, usually through automated pipelines. Even a GSP may give a misleading assignment when used as a reference: the homolog may be close enough to support isofunctionality, but the substrate of the GSP is absent from the species being annotated. In such cases, the enzymes cannot be isofunctional. Here, we examined a variety of such issues in halophilic archaea (class Halobacteria), with a strong focus on the model haloarchaeon Haloferax volcanii.
Results: Annotated proteins of Hfx. volcanii were identified for which public databases tend to assign a function that is probably incorrect. In some cases, an alternative, probably correct, function can be predicted or inferred from the available evidence, but this has not been adopted by public databases because experimental validation is lacking. In other cases, a probably invalid specific function is predicted by homology, and while there is evidence that this assigned function is unlikely, the true function remains elusive. We listed 50 of those cases, each with detailed background information, so that a conclusion about the most likely biological function can be drawn. For reasons of brevity and comprehension, only the key aspects are listed in the main text, with detailed information being provided in a corresponding section of the Supplementary Materials.
Conclusions: Compiling, describing and summarizing these open annotation issues and functional predictions will benefit the scientific community in the general effort to improve the evaluation of protein function assignments and more thoroughly detail them. By highlighting the gaps and likely annotation errors currently in the databases, we hope this study will provide a framework for experimentalists to systematically confirm (or disprove) our function predictions or to uncover yet more unexpected functions.
Keywords: Gold Standard Protein; Haloferax volcanii; annotation error; genome annotation; haloarchaea.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

Similar articles
-
Towards glycoengineering in archaea: replacement of Haloferax volcanii AglD with homologous glycosyltransferases from other halophilic archaea.Appl Environ Microbiol. 2010 Sep;76(17):5684-92. doi: 10.1128/AEM.00681-10. Epub 2010 Jul 2. Appl Environ Microbiol. 2010. PMID: 20601508 Free PMC article.
-
Overexpression and purification of halophilic proteins in Haloferax volcanii.Bioeng Bugs. 2010 Jul-Aug;1(4):288-90. doi: 10.4161/bbug.1.4.11794. Epub 2010 Mar 17. Bioeng Bugs. 2010. PMID: 21327063 Free PMC article.
-
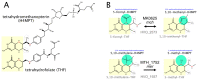
Conserved active site cysteine residue of archaeal THI4 homolog is essential for thiamine biosynthesis in Haloferax volcanii.BMC Microbiol. 2014 Oct 28;14:260. doi: 10.1186/s12866-014-0260-0. BMC Microbiol. 2014. PMID: 25348237 Free PMC article.
-
Post-translation modification in Archaea: lessons from Haloferax volcanii and other haloarchaea.FEMS Microbiol Rev. 2013 Jul;37(4):583-606. doi: 10.1111/1574-6976.12012. Epub 2012 Dec 20. FEMS Microbiol Rev. 2013. PMID: 23167813 Free PMC article. Review.
-
An Experimental Approach to Genome Annotation: This report is based on a colloquium sponsored by the American Academy of Microbiology held July 19-20, 2004, in Washington, DC.Washington (DC): American Society for Microbiology; 2004. Washington (DC): American Society for Microbiology; 2004. PMID: 33001599 Free Books & Documents. Review.
Cited by
-
Diversity, taxonomy, and evolution of archaeal viruses of the class Caudoviricetes.PLoS Biol. 2021 Nov 9;19(11):e3001442. doi: 10.1371/journal.pbio.3001442. eCollection 2021 Nov. PLoS Biol. 2021. PMID: 34752450 Free PMC article.
-
A crowdsourcing open platform for literature curation in UniProt.PLoS Biol. 2021 Dec 6;19(12):e3001464. doi: 10.1371/journal.pbio.3001464. eCollection 2021 Dec. PLoS Biol. 2021. PMID: 34871295 Free PMC article.
-
Comparative Analysis of rRNA Removal Methods for RNA-Seq Differential Expression in Halophilic Archaea.Biomolecules. 2022 May 10;12(5):682. doi: 10.3390/biom12050682. Biomolecules. 2022. PMID: 35625610 Free PMC article.
References
-
- Schulze S., Adams Z., Cerletti M., De Castro R., Ferreira-Cerca S., Fufezan C., Gimenez M.I., Hippler M., Jevtic Z., Knuppel R., et al. The Archaeal Proteome Project advances knowledge about archaeal cell biology through comprehensive proteomics. Nat. Commun. 2020;11:3145. doi: 10.1038/s41467-020-16784-7. - DOI - PMC - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

