miR-375 prevents high-fat diet-induced insulin resistance and obesity by targeting the aryl hydrocarbon receptor and bacterial tryptophanase (tnaA) gene
- PMID: 33754048
- PMCID: PMC7977461
- DOI: 10.7150/thno.52558
miR-375 prevents high-fat diet-induced insulin resistance and obesity by targeting the aryl hydrocarbon receptor and bacterial tryptophanase (tnaA) gene
Abstract
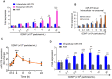
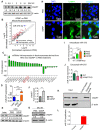
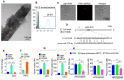
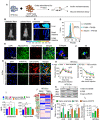
Background: Diet manipulation is the basis for prevention of obesity and diabetes. The molecular mechanisms that mediate the diet-based prevention of insulin resistance are not well understood. Here, as proof-of-concept, ginger-derived nanoparticles (GDNP) were used for studying molecular mechanisms underlying GDNP mediated prevention of high-fat diet induced insulin resistance. Methods: Ginger-derived nanoparticles (GDNP) were isolated from ginger roots and administered orally to C57BL/6 high-fat diet mice. Fecal exosomes released from intestinal epithelial cells (IECs) of PBS or GDNP treated high-fat diet (HFD) fed mice were isolated by differential centrifugation. A micro-RNA (miRNA) polymerase chain reaction (PCR) array was used to profile the exosomal miRs and miRs of interest were further analyzed by quantitative real time (RT) PCR. miR-375 or antisense-miR375 was packed into nanoparticles made from the lipids extracted from GDNP. Nanoparticles was fluorescent labeled for monitoring their in vivo trafficking route after oral administration. The effect of these nanoparticles on glucose and insulin response of mice was determined by glucose and insulin tolerance tests. Results: We report that HFD feeding increased the expression of AhR and inhibited the expression of miR-375 and VAMP7. Treatment with orally administered ginger-derived nanoparticles (GDNP) resulted in reversing HFD mediated inhibition of the expression of miR-375 and VAMP7. miR-375 knockout mice exhibited impaired glucose homeostasis and insulin resistance. Induction of intracellular miR-375 led to inhibition of the expression of AhR and VAMP7 mediated exporting of miR-375 into intestinal epithelial exosomes where they were taken up by gut bacteria and inhibited the production of the AhR ligand indole. Intestinal exosomes can also traffic to the liver and be taken up by hepatocytes, leading to miR-375 mediated inhibition of hepatic AhR over-expression and inducing the expression of genes associated with the hepatic insulin response. Altogether, GDNP prevents high-fat diet-induced insulin resistance by miR-375 mediated inhibition of the aryl hydrocarbon receptor mediated pathways over activated by HFD feeding. Conclusion: Collectively our findings reveal that oral administration of GDNP to HFD mice improves host glucose tolerance and insulin response via regulating AhR expression by GDNP induced miR-375 and VAMP7.
Keywords: AhR; E. coli tryptophanase (tnaA); Exosomes; Ginger derived nanoparticles; VAMP7; gut/liver axis; indole; insulin resistance.; miR-375.
© The author(s).
Conflict of interest statement
Competing Interests: The authors have declared that no competing interest exists.
Figures


Comment in
-
MicroRNA loaded edible nanoparticles: an emerging personalized therapeutic approach for the treatment of obesity and metabolic disorders.Theranostics. 2022 Mar 5;12(6):2631-2634. doi: 10.7150/thno.71399. eCollection 2022. Theranostics. 2022. PMID: 35401814 Free PMC article.
Similar articles
-
Ginger nanoparticles mediated induction of Foxa2 prevents high-fat diet-induced insulin resistance.Theranostics. 2022 Jan 1;12(3):1388-1403. doi: 10.7150/thno.62514. eCollection 2022. Theranostics. 2022. PMID: 35154496 Free PMC article.
-
Garlic exosome-like nanoparticles reverse high-fat diet induced obesity via the gut/brain axis.Theranostics. 2022 Jan 1;12(3):1220-1246. doi: 10.7150/thno.65427. eCollection 2022. Theranostics. 2022. PMID: 35154484 Free PMC article.
-
Pdgfrα-Cre mediated knockout of the aryl hydrocarbon receptor protects mice from high-fat diet induced obesity and hepatic steatosis.PLoS One. 2020 Jul 30;15(7):e0236741. doi: 10.1371/journal.pone.0236741. eCollection 2020. PLoS One. 2020. PMID: 32730300 Free PMC article.
-
High-fat diet-induced upregulation of exosomal phosphatidylcholine contributes to insulin resistance.Nat Commun. 2021 Jan 11;12(1):213. doi: 10.1038/s41467-020-20500-w. Nat Commun. 2021. PMID: 33431899 Free PMC article.
-
Metabolic Side Effects from Antipsychotic Treatment with Clozapine Linked to Aryl Hydrocarbon Receptor (AhR) Activation.Biomedicines. 2024 Oct 10;12(10):2294. doi: 10.3390/biomedicines12102294. Biomedicines. 2024. PMID: 39457607 Free PMC article. Review.
Cited by
-
Ginger: a representative material of herb-derived exosome-like nanoparticles.Front Nutr. 2023 Jul 13;10:1223349. doi: 10.3389/fnut.2023.1223349. eCollection 2023. Front Nutr. 2023. PMID: 37521414 Free PMC article. Review.
-
Emerging Microfluidic Tools for Simultaneous Exosomes and Cargo Biosensing in Liquid Biopsy: New Integrated Miniaturized FFF-Assisted Approach for Colon Cancer Diagnosis.Sensors (Basel). 2023 Nov 27;23(23):9432. doi: 10.3390/s23239432. Sensors (Basel). 2023. PMID: 38067805 Free PMC article. Review.
-
Fatty Exosomes Aggravate Metabolic Disorders.Mol Cells. 2022 Oct 31;45(10):692-694. doi: 10.14348/molcells.2022.0135. Epub 2022 Oct 6. Mol Cells. 2022. PMID: 36254711 Free PMC article. No abstract available.
-
MicroRNAs as Epigenetic Regulators of Obesity.Adv Exp Med Biol. 2024;1460:595-627. doi: 10.1007/978-3-031-63657-8_20. Adv Exp Med Biol. 2024. PMID: 39287866 Review.
-
New insights into the role of dietary triglyceride absorption in obesity and metabolic diseases.Front Pharmacol. 2023 Feb 2;14:1097835. doi: 10.3389/fphar.2023.1097835. eCollection 2023. Front Pharmacol. 2023. PMID: 36817150 Free PMC article. Review.
References
-
- Rothhammer V, Quintana FJ. The aryl hydrocarbon receptor: an environmental sensor integrating immune responses in health and disease. Nature reviews Immunology. 2019;19:184–97. - PubMed
-
- Roh E, Kwak SH, Jung HS, Cho YM, Pak YK, Park KS. et al. Serum aryl hydrocarbon receptor ligand activity is associated with insulin resistance and resulting type 2 diabetes. Acta Diabetol. 2015;52:489–95. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

