How Rhizobia Adapt to the Nodule Environment
- PMID: 33526611
- PMCID: PMC8315917
- DOI: 10.1128/JB.00539-20
How Rhizobia Adapt to the Nodule Environment
Abstract
Rhizobia are a phylogenetically diverse group of soil bacteria that engage in mutualistic interactions with legume plants. Although specifics of the symbioses differ between strains and plants, all symbioses ultimately result in the formation of specialized root nodule organs that host the nitrogen-fixing microsymbionts called bacteroids. Inside nodules, bacteroids encounter unique conditions that necessitate the global reprogramming of physiological processes and the rerouting of their metabolism. Decades of research have addressed these questions using genetics, omics approaches, and, more recently, computational modeling. Here, we discuss the common adaptations of rhizobia to the nodule environment that define the core principles of bacteroid functioning. All bacteroids are growth arrested and perform energy-intensive nitrogen fixation fueled by plant-provided C4-dicarboxylates at nanomolar oxygen levels. At the same time, bacteroids are subject to host control and sanctioning that ultimately determine their fitness and have fundamental importance for the evolution of a stable mutualistic relationship.
Keywords: growth arrest; host sanctioning; metabolism; microaerobiosis; modeling; nitrogen fixation; rhizobium-legume symbiosis; root nodule.
Figures
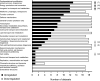
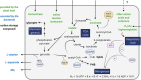
Similar articles
-
Proteomic analysis dissects the impact of nodulation and biological nitrogen fixation on Vicia faba root nodule physiology.Plant Mol Biol. 2018 Jun;97(3):233-251. doi: 10.1007/s11103-018-0736-7. Epub 2018 May 19. Plant Mol Biol. 2018. PMID: 29779088
-
Starting points in plant-bacteria nitrogen-fixing symbioses: intercellular invasion of the roots.J Exp Bot. 2017 Apr 1;68(8):1905-1918. doi: 10.1093/jxb/erw387. J Exp Bot. 2017. PMID: 27756807 Review.
-
A Minimal Genetic Passkey to Unlock Many Legume Doors to Root Nodulation by Rhizobia.Genes (Basel). 2020 May 7;11(5):521. doi: 10.3390/genes11050521. Genes (Basel). 2020. PMID: 32392829 Free PMC article.
-
From Intracellular Bacteria to Differentiated Bacteroids: Transcriptome and Metabolome Analysis in Aeschynomene Nodules Using the Bradyrhizobium sp. Strain ORS285 bclA Mutant.J Bacteriol. 2019 Aug 8;201(17):e00191-19. doi: 10.1128/JB.00191-19. Print 2019 Sep 1. J Bacteriol. 2019. PMID: 31182497 Free PMC article.
-
What determines the efficiency of N(2)-fixing Rhizobium-legume symbioses?Adv Microb Physiol. 2012;60:325-89. doi: 10.1016/B978-0-12-398264-3.00005-X. Adv Microb Physiol. 2012. PMID: 22633062 Review.
Cited by
-
Bradyrhizobium diazoefficiens Requires Chemical Chaperones To Cope with Osmotic Stress during Soybean Infection.mBio. 2021 Mar 30;12(2):e00390-21. doi: 10.1128/mBio.00390-21. mBio. 2021. PMID: 33785618 Free PMC article.
-
Sinorhizobium meliloti DnaJ Is Required for Surface Motility, Stress Tolerance, and for Efficient Nodulation and Symbiotic Nitrogen Fixation.Int J Mol Sci. 2023 Mar 19;24(6):5848. doi: 10.3390/ijms24065848. Int J Mol Sci. 2023. PMID: 36982921 Free PMC article.
-
Interaction between bacterial endophytes and host plants.Front Plant Sci. 2023 Jan 18;13:1092105. doi: 10.3389/fpls.2022.1092105. eCollection 2022. Front Plant Sci. 2023. PMID: 36743537 Free PMC article. Review.
-
Competition, Nodule Occupancy, and Persistence of Inoculant Strains: Key Factors in the Rhizobium-Legume Symbioses.Front Plant Sci. 2021 Aug 19;12:690567. doi: 10.3389/fpls.2021.690567. eCollection 2021. Front Plant Sci. 2021. PMID: 34489993 Free PMC article. Review.
-
Metagenomic analysis of Raphidiopsisraciborskii microbiome: beyond the individual.Biodivers Data J. 2021 Oct 21;9:e72514. doi: 10.3897/BDJ.9.e72514. eCollection 2021. Biodivers Data J. 2021. PMID: 34754266 Free PMC article.
References
-
- Radhakrishnan GV, Keller J, Rich MK, Vernie T, Mbadinga Mbadinga DL, Vigneron N, Cottret L, Clemente HS, Libourel C, Cheema J, Linde A-M, Eklund DM, Cheng S, Wong GKS, Lagercrantz U, Li F-W, Oldroyd GED, Delaux P-M. 2020. An ancestral signalling pathway is conserved in intracellular symbioses-forming plant lineages. Nat Plants 6:280–289. 10.1038/s41477-020-0613-7. - DOI - PubMed
-
- Gherbi H, Markmann K, Svistoonoff S, Estevan J, Autran D, Giczey G, Auguy F, Peret B, Laplaze L, Franche C, Parniske M, Bogusz D. 2008. SymRK defines a common genetic basis for plant root endosymbioses with arbuscular mycorrhiza fungi, rhizobia, and Frankia bacteria. Proc Natl Acad Sci U S A 105:4928–4932. 10.1073/pnas.0710618105. - DOI - PMC - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

