Genome mining and UHPLC-QTOF-MS/MS to identify the potential antimicrobial compounds and determine the specificity of biosynthetic gene clusters in Bacillus subtilis NCD-2
- PMID: 33153447
- PMCID: PMC7643408
- DOI: 10.1186/s12864-020-07160-2
Genome mining and UHPLC-QTOF-MS/MS to identify the potential antimicrobial compounds and determine the specificity of biosynthetic gene clusters in Bacillus subtilis NCD-2
Abstract
Background: Bacillus subtilis strain NCD-2 is an excellent biocontrol agent against plant soil-borne diseases and shows broad-spectrum antifungal activities. This study aimed to explore some secondary metabolite biosynthetic gene clusters and related antimicrobial compounds in strain NCD-2. An integrative approach combining genome mining and structural identification technologies using ultra-high-performance liquid chromatography coupled to quadrupole time-of-flight tandem mass spectrometry (UHPLC-MS/MS), was adopted to interpret the chemical origins of metabolites with significant biological activities.
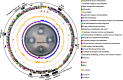
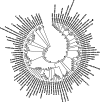
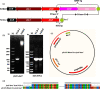
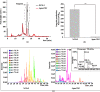
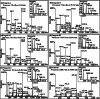
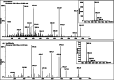
Results: Genome mining revealed nine gene clusters encoding secondary metabolites with predicted functions, including fengycin, surfactin, bacillaene, subtilosin, bacillibactin, bacilysin and three unknown products. Fengycin, surfactin, bacillaene and bacillibactin were successfully detected from the fermentation broth of strain NCD-2 by UHPLC-QTOF-MS/MS. The biosynthetic gene clusters of bacillaene, subtilosin, bacillibactin, and bacilysin showed 100% amino acid sequence identities with those in B. velezensis strain FZB42, whereas the identities of the surfactin and fengycin gene clusters were only 83 and 92%, respectively. Further comparison revealed that strain NCD-2 had lost the fenC and fenD genes in the fengycin biosynthetic operon. The biosynthetic enzyme-related gene srfAB for surfactin was divided into two parts. Bioinformatics analysis suggested that FenE in strain NCD-2 had a similar function to FenE and FenC in strain FZB42, and that FenA in strain NCD-2 had a similar function to FenA and FenD in strain FZB42. Five different kinds of fengycins, with 26 homologs, and surfactin, with 4 homologs, were detected from strain NCD-2. To the best of our knowledge, this is the first report of a non-typical gene cluster related to fengycin synthesis.
Conclusions: Our study revealed a number of gene clusters encoding antimicrobial compounds in the genome of strain NCD-2, including a fengycin synthetic gene cluster that might be unique by using genome mining and UHPLC-QTOF-MS/MS. The production of fengycin, surfactin, bacillaene and bacillibactin might explain the biological activities of strain NCD-2.
Keywords: Bacillus subtilis NCD-2; Fengycin; Genome mining; Secondary metabolites; UHPLC–QTOF–MS/MS.
Conflict of interest statement
The authors declare that they have no competing interests.
Figures


Similar articles
-
Genomic and metabolomic insights into the antimicrobial compounds and plant growth-promoting potential of Bacillus velezensis Q-426.BMC Genomics. 2023 Oct 4;24(1):589. doi: 10.1186/s12864-023-09662-1. BMC Genomics. 2023. PMID: 37794314 Free PMC article.
-
A comprehensive understanding of the biocontrol potential of Bacillus velezensis LM2303 against Fusarium head blight.PLoS One. 2018 Jun 1;13(6):e0198560. doi: 10.1371/journal.pone.0198560. eCollection 2018. PLoS One. 2018. PMID: 29856856 Free PMC article.
-
Comparative Genome Analysis Reveals Phylogenetic Identity of Bacillus velezensis HNA3 and Genomic Insights into Its Plant Growth Promotion and Biocontrol Effects.Microbiol Spectr. 2022 Feb 23;10(1):e0216921. doi: 10.1128/spectrum.02169-21. Epub 2022 Feb 2. Microbiol Spectr. 2022. PMID: 35107331 Free PMC article.
-
Bacillus velezensis: A Valuable Member of Bioactive Molecules within Plant Microbiomes.Molecules. 2019 Mar 16;24(6):1046. doi: 10.3390/molecules24061046. Molecules. 2019. PMID: 30884857 Free PMC article. Review.
-
Biological control of plant pathogens by Bacillus species.J Biotechnol. 2018 Nov 10;285:44-55. doi: 10.1016/j.jbiotec.2018.07.044. Epub 2018 Aug 30. J Biotechnol. 2018. PMID: 30172784 Review.
Cited by
-
Evaluation and Genome Analysis of Bacillus subtilis YB-04 as a Potential Biocontrol Agent Against Fusarium Wilt and Growth Promotion Agent of Cucumber.Front Microbiol. 2022 Jun 9;13:885430. doi: 10.3389/fmicb.2022.885430. eCollection 2022. Front Microbiol. 2022. PMID: 35756052 Free PMC article.
-
A Metagenomic Nanopore Sequence Analysis Combined with Conventional Screening and Spectroscopic Methods for Deciphering the Antimicrobial Metabolites Produced by Alcaligenes faecalis Soil Isolate MZ921504.Antibiotics (Basel). 2021 Nov 11;10(11):1382. doi: 10.3390/antibiotics10111382. Antibiotics (Basel). 2021. PMID: 34827320 Free PMC article.
-
Characterization and genome mining of Bacillus subtilis BDSA1 isolated from river water in Bangladesh: A promising bacterium with diverse biotechnological applications.Heliyon. 2024 Jul 11;10(14):e34369. doi: 10.1016/j.heliyon.2024.e34369. eCollection 2024 Jul 30. Heliyon. 2024. PMID: 39114027 Free PMC article.
-
Colonization Ability of Bacillus subtilis NCD-2 in Different Crops and Its Effect on Rhizosphere Microorganisms.Microorganisms. 2023 Mar 17;11(3):776. doi: 10.3390/microorganisms11030776. Microorganisms. 2023. PMID: 36985349 Free PMC article.
-
Genomic insights into Bacillus subtilis MBB3B9 mediated aluminium stress mitigation for enhanced rice growth.Sci Rep. 2023 Sep 30;13(1):16467. doi: 10.1038/s41598-023-42804-9. Sci Rep. 2023. PMID: 37777563 Free PMC article.
References
-
- Wang P, Guo Q, Ma Y, Li S, Lu X, Zhang X, Ma P. DegQ regulates the production of fengycins and biofilm formation of the biocontrol agent Bacillus subtilis NCD-2. Microbiol Res. 2015;178:42–50. - PubMed
-
- Fan H, Ru J, Zhang Y, Wang Q, Li Y. Fengycin produced by Bacillus subtilis 9407 plays a major role in the biocontrol of apple ring rot disease. Microbiol Res. 2017;199:89–97. - PubMed
-
- Sonenshein AL. Control of sporulation initiation in Bacillus subtilis. Curr Opin Microbiol. 2000;3(6):561–566. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

