Lysogeny in the Lactic Acid Bacterium Oenococcus oeni Is Responsible for Modified Colony Morphology on Red Grape Juice Agar
- PMID: 31375489
- PMCID: PMC6752011
- DOI: 10.1128/AEM.00997-19
Lysogeny in the Lactic Acid Bacterium Oenococcus oeni Is Responsible for Modified Colony Morphology on Red Grape Juice Agar
Abstract
Oenococcus oeni is the lactic acid bacterium (LAB) that most commonly drives malolactic fermentation in wine. Although oenococcal prophages are highly prevalent, their implications on bacterial fitness have remained unexplored and more research is required in this field. An important step toward achieving this goal is the ability to produce isogenic pairs of strains that differ only by the lysogenic presence of a given prophage, allowing further comparisons of different phenotypic traits. A novel protocol for the rapid isolation of lysogens is presented. Bacteria were first picked from the center of turbid plaques produced by temperate oenophages on a sensitive nonlysogenic host. When streaked onto an agar medium containing red grape juice (RGJ), cells segregated into white and red colonies. PCR amplifications with phage-specific primers demonstrated that only lysogens underwent white-red morphotypic switching. The method proved successful for various oenophages irrespective of their genomic content and attachment site used for site-specific recombination in the bacterial chromosome. The color switch was also observed when a sensitive nonlysogenic strain was infected with an exogenously provided lytic phage, suggesting that intracolonial lysis triggers the change. Last, lysogens also produced red colonies on white grape juice agar supplemented with polyphenolic compounds. We posit that spontaneous prophage excision produces cell lysis events in lysogenic colonies growing on RGJ agar, which, in turn, foster interactions between lysed materials and polyphenolic compounds to yield colonies easily distinguishable by their red color. Furthermore, the technique was used successfully with other species of LAB.IMPORTANCE The presence of white and red colonies on red grape juice (RGJ) agar during enumeration of Oenococcus oeni in wine samples is frequently observed by stakeholders in the wine industry. Our study brings an explanation for this intriguing phenomenon and establishes a link between the white-red color switch and the lysogenic state of O. oeni It also provides a simple and inexpensive method to distinguish between lysogenic and nonlysogenic derivatives in O. oeni with a minimum of expended time and effort. Noteworthy, the protocol could be adapted to two other species of LAB, namely, Leuconostoc citreum and Lactobacillus plantarum It could be an effective tool to provide genetic, ecological, and functional insights into lysogeny and aid in improving biotechnological processes involving members of the lactic acid bacterium (LAB) family.
Keywords: bacteriophages; lactic acid bacteria; lysogeny.
Copyright © 2019 American Society for Microbiology.
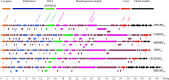
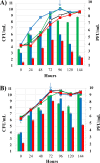
Figures





Similar articles
-
Assessment of the lysogenic status in the lactic acid bacterium O. oeni during the spontaneous malolactic fermentation of red wines.Food Microbiol. 2022 May;103:103947. doi: 10.1016/j.fm.2021.103947. Epub 2021 Nov 16. Food Microbiol. 2022. PMID: 35082064
-
Expanding the diversity of oenococcal bacteriophages: insights into a novel group based on the integrase sequence.Int J Food Microbiol. 2013 Sep 2;166(2):331-40. doi: 10.1016/j.ijfoodmicro.2013.06.032. Epub 2013 Jul 7. Int J Food Microbiol. 2013. PMID: 23994162
-
Oenococcus oeni and the genomic era.FEMS Microbiol Rev. 2017 Aug 1;41(Supp_1):S84-S94. doi: 10.1093/femsre/fux034. FEMS Microbiol Rev. 2017. PMID: 28830095 Review.
-
Development of a new method for detection and identification of Oenococcus oeni bacteriophages based on endolysin gene sequence and randomly amplified polymorphic DNA.Appl Environ Microbiol. 2013 Aug;79(16):4799-805. doi: 10.1128/AEM.01307-13. Epub 2013 May 31. Appl Environ Microbiol. 2013. PMID: 23728816 Free PMC article.
-
Distribution of Oenococcus oeni populations in natural habitats.Appl Microbiol Biotechnol. 2019 Apr;103(7):2937-2945. doi: 10.1007/s00253-019-09689-z. Epub 2019 Feb 20. Appl Microbiol Biotechnol. 2019. PMID: 30788540 Free PMC article. Review.
Cited by
-
Characterization of the First Virulent Phage Infecting Oenococcus oeni, the Queen of the Cellars.Front Microbiol. 2021 Jan 13;11:596541. doi: 10.3389/fmicb.2020.596541. eCollection 2020. Front Microbiol. 2021. PMID: 33519734 Free PMC article.
-
Draft Genome Sequence of Oenococcus kitaharae CRBO2176, Isolated from Homemade Water Kefir.Microbiol Resour Announc. 2023 Apr 18;12(4):e0107222. doi: 10.1128/mra.01072-22. Epub 2023 Mar 29. Microbiol Resour Announc. 2023. PMID: 36988513 Free PMC article.
-
"French Phage Network" Annual Conference-Fifth Meeting Report.Viruses. 2020 Apr 14;12(4):446. doi: 10.3390/v12040446. Viruses. 2020. PMID: 32295276 Free PMC article.
References
Publication types
MeSH terms
Substances
Supplementary concepts
LinkOut - more resources
Full Text Sources
Miscellaneous

