A microfluidic device for inferring metabolic landscapes in yeast monolayer colonies
- PMID: 31259688
- PMCID: PMC6624017
- DOI: 10.7554/eLife.47951
A microfluidic device for inferring metabolic landscapes in yeast monolayer colonies
Abstract
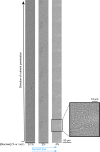
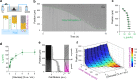
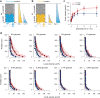
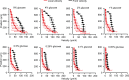
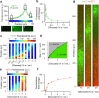
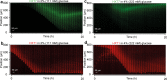
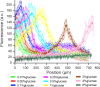
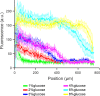
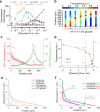
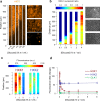
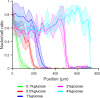
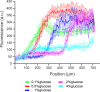
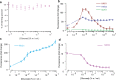
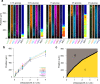
Microbial colonies are fascinating structures in which growth and internal organization reflect complex morphogenetic processes. Here, we generated a microfluidics device with arrays of long monolayer yeast colonies to further global understanding of how intercellular metabolic interactions affect the internal structure of colonies within defined boundary conditions. We observed the emergence of stable glucose gradients using fluorescently labeled hexose transporters and quantified the spatial correlations with intra-colony growth rates and expression of other genes regulated by glucose availability. These landscapes depended on the external glucose concentration as well as secondary gradients, for example amino acid availability. This work demonstrates the regulatory genetic networks governing cellular physiological adaptation are the key to internal structuration of cellular assemblies. This approach could be used in the future to decipher the interplay between long-range metabolic interactions, cellular development and morphogenesis in more complex systems.
Keywords: S. cerevisiae; gene expression; metabolism; microfluidics; physics of living systems; spatial organization; yeast colony.
© 2019, Marinkovic et al.
Conflict of interest statement
ZM, CV, MA, XS, JD, AL, PH No competing interests declared
Figures




Similar articles
-
Observing Nutrient Gradients, Gene Expression and Growth Variation Using the "Yeast Machine" Microfluidic Device.Bio Protoc. 2020 Jul 5;10(13):e3668. doi: 10.21769/BioProtoc.3668. eCollection 2020 Jul 5. Bio Protoc. 2020. PMID: 33659338 Free PMC article.
-
Saccharomyces cerevisiae and metabolic activators: HXT3 gene expression and fructose/glucose discrepancy in sluggish fermentation conditions.World J Microbiol Biotechnol. 2016 Dec;32(12):196. doi: 10.1007/s11274-016-2154-9. Epub 2016 Oct 12. World J Microbiol Biotechnol. 2016. PMID: 27734279
-
Growing yeast into cylindrical colonies.Biophys J. 2014 May 20;106(10):2214-21. doi: 10.1016/j.bpj.2014.02.040. Biophys J. 2014. PMID: 24853750 Free PMC article.
-
Glucose- and nitrogen sensing and regulatory mechanisms in Saccharomyces cerevisiae.FEMS Yeast Res. 2014 Aug;14(5):683-96. doi: 10.1111/1567-1364.12157. Epub 2014 May 8. FEMS Yeast Res. 2014. PMID: 24738657 Review.
-
The glucose signaling network in yeast.Biochim Biophys Acta. 2013 Nov;1830(11):5204-10. doi: 10.1016/j.bbagen.2013.07.025. Epub 2013 Aug 2. Biochim Biophys Acta. 2013. PMID: 23911748 Free PMC article. Review.
Cited by
-
Preventing Photomorbidity in Long-Term Multi-color Fluorescence Imaging of Saccharomyces cerevisiae and S. pombe.G3 (Bethesda). 2020 Dec 3;10(12):4373-4385. doi: 10.1534/g3.120.401465. G3 (Bethesda). 2020. PMID: 33023973 Free PMC article.
-
Aspects of Multicellularity in Saccharomyces cerevisiae Yeast: A Review of Evolutionary and Physiological Mechanisms.Genes (Basel). 2020 Jun 24;11(6):690. doi: 10.3390/genes11060690. Genes (Basel). 2020. PMID: 32599749 Free PMC article. Review.
-
Chemotropism among populations of yeast cells with spatiotemporal resolution in a biofabricated microfluidic platform.Biomicrofluidics. 2020 Jan 17;14(1):014108. doi: 10.1063/1.5128739. eCollection 2020 Jan. Biomicrofluidics. 2020. PMID: 32002107 Free PMC article.
-
CyberSco.Py an open-source software for event-based, conditional microscopy.Sci Rep. 2022 Jul 8;12(1):11579. doi: 10.1038/s41598-022-15207-5. Sci Rep. 2022. PMID: 35803978 Free PMC article.
-
Diverse conditions support near-zero growth in yeast: Implications for the study of cell lifespan.Microb Cell. 2019 Aug 20;6(9):397-413. doi: 10.15698/mic2019.09.690. Microb Cell. 2019. PMID: 31528631 Free PMC article. Review.
References
-
- Antwis RE, Griffiths SM, Harrison XA, Aranega-Bou P, Arce A, Bettridge AS, Brailsford FL, de Menezes A, Devaynes A, Forbes KM, Fry EL, Goodhead I, Haskell E, Heys C, James C, Johnston SR, Lewis GR, Lewis Z, Macey MC, McCarthy A, McDonald JE, Mejia-Florez NL, O’Brien D, Orland C, Pautasso M, Reid WDK, Robinson HA, Wilson K, Sutherland WJ. Fifty important research questions in microbial ecology. FEMS Microbiology Ecology. 2017;93:fix044. doi: 10.1093/femsec/fix044. - DOI - PubMed
-
- Bonander N, Ferndahl C, Mostad P, Wilks MD, Chang C, Showe L, Gustafsson L, Larsson C, Bill RM. Transcriptome analysis of a respiratory Saccharomyces cerevisiae strain suggests the expression of its phenotype is glucose insensitive and predominantly controlled by Hap4, Cat8 and Mig1. BMC Genomics. 2008;9:365. doi: 10.1186/1471-2164-9-365. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- 724813/European Commission/International
- ICEBERG-ANR-10-BINF-06-01/Agence Nationale de la Recherche/International
- ANR-16-CE12-0025-01/Agence Nationale de la Recherche/International
- ANR-11-LABX-0071/Agence Nationale de la Recherche/International
- ANR-11-IDEX-0005-01/Agence Nationale de la Recherche/International
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

