Homozygous frameshift mutations in FAT1 cause a syndrome characterized by colobomatous-microphthalmia, ptosis, nephropathy and syndactyly
- PMID: 30862798
- PMCID: PMC6414540
- DOI: 10.1038/s41467-019-08547-w
Homozygous frameshift mutations in FAT1 cause a syndrome characterized by colobomatous-microphthalmia, ptosis, nephropathy and syndactyly
Abstract
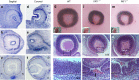
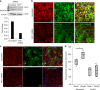
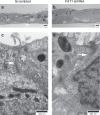
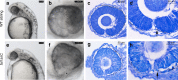
A failure in optic fissure fusion during development can lead to blinding malformations of the eye. Here, we report a syndrome characterized by facial dysmorphism, colobomatous microphthalmia, ptosis and syndactyly with or without nephropathy, associated with homozygous frameshift mutations in FAT1. We show that Fat1 knockout mice and zebrafish embryos homozygous for truncating fat1a mutations exhibit completely penetrant coloboma, recapitulating the most consistent developmental defect observed in affected individuals. In human retinal pigment epithelium (RPE) cells, the primary site for the fusion of optic fissure margins, FAT1 is localized at earliest cell-cell junctions, consistent with a role in facilitating optic fissure fusion during vertebrate eye development. Our findings establish FAT1 as a gene with pleiotropic effects in human, in that frameshift mutations cause a severe multi-system disorder whereas recessive missense mutations had been previously associated with isolated glomerulotubular nephropathy.
Conflict of interest statement
The authors declare no competing interests.
Figures

Similar articles
-
Identification of causative gene mutation in an Iranian family with coloboma and nephropathy using whole exome sequencing.CEN Case Rep. 2022 Nov;11(4):404-407. doi: 10.1007/s13730-022-00692-4. Epub 2022 Feb 18. CEN Case Rep. 2022. PMID: 35179696 Free PMC article.
-
ABCB6 mutations cause ocular coloboma.Am J Hum Genet. 2012 Jan 13;90(1):40-8. doi: 10.1016/j.ajhg.2011.11.026. Epub 2012 Jan 5. Am J Hum Genet. 2012. PMID: 22226084 Free PMC article.
-
Mutations in c12orf57 cause a syndromic form of colobomatous microphthalmia.Am J Hum Genet. 2013 Mar 7;92(3):387-91. doi: 10.1016/j.ajhg.2013.01.008. Epub 2013 Feb 28. Am J Hum Genet. 2013. PMID: 23453665 Free PMC article.
-
Ocular coloboma: Genetic variants reveal a dynamic model of eye development.Am J Med Genet C Semin Med Genet. 2020 Sep;184(3):590-610. doi: 10.1002/ajmg.c.31831. Epub 2020 Aug 27. Am J Med Genet C Semin Med Genet. 2020. PMID: 32852110 Review.
-
Genetics of microphthalmos.Int Ophthalmol. 1981 Aug;4(1-2):45-65. doi: 10.1007/BF00139580. Int Ophthalmol. 1981. PMID: 6795139 Review.
Cited by
-
Expanding the Spectrum of FAT1 Nephropathies by Novel Mutations That Affect Hippo Signaling.Kidney Int Rep. 2021 Jan 29;6(5):1368-1378. doi: 10.1016/j.ekir.2021.01.023. eCollection 2021 May. Kidney Int Rep. 2021. PMID: 34013115 Free PMC article.
-
Nf2 fine-tunes proliferation and tissue alignment during closure of the optic fissure in the embryonic mouse eye.Hum Mol Genet. 2020 Dec 18;29(20):3373-3387. doi: 10.1093/hmg/ddaa228. Hum Mol Genet. 2020. PMID: 33075808 Free PMC article.
-
FAT1 expression in T-cell acute lymphoblastic leukemia (T-ALL) modulates proliferation and WNT signaling.Sci Rep. 2023 Jan 18;13(1):972. doi: 10.1038/s41598-023-27792-0. Sci Rep. 2023. PMID: 36653435 Free PMC article.
-
Whole Exome Sequencing Is the Minimal Technological Approach in Probands Born to Consanguineous Couples.Genes (Basel). 2021 Jun 24;12(7):962. doi: 10.3390/genes12070962. Genes (Basel). 2021. PMID: 34202629 Free PMC article.
-
Genetic heterogeneity in familial forms of genetic generalized epilepsy: from mono- to oligogenism.Hum Genomics. 2024 Nov 21;18(1):130. doi: 10.1186/s40246-024-00659-9. Hum Genomics. 2024. PMID: 39574152 Free PMC article.
References
-
- Patel, A. & Sowden, J. C. Genes and pathways in optic fissure closure. Semin. Cell. Dev. Biol. 10.1016/j.semcdb.2017.10.010 (2017). - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

