Mapping the Physiological Response of Oenococcus oeni to Ethanol Stress Using an Extended Genome-Scale Metabolic Model
- PMID: 29545779
- PMCID: PMC5838312
- DOI: 10.3389/fmicb.2018.00291
Mapping the Physiological Response of Oenococcus oeni to Ethanol Stress Using an Extended Genome-Scale Metabolic Model
Abstract
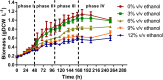
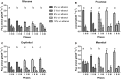
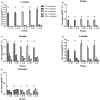
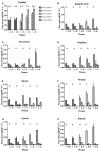
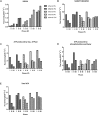
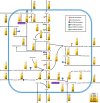
The effect of ethanol on the metabolism of Oenococcus oeni, the bacterium responsible for the malolactic fermentation (MLF) of wine, is still scarcely understood. Here, we characterized the global metabolic response in O. oeni PSU-1 to increasing ethanol contents, ranging from 0 to 12% (v/v). We first optimized a wine-like, defined culture medium, MaxOeno, to allow sufficient bacterial growth to be able to quantitate different metabolites in batch cultures of O. oeni. Then, taking advantage of the recently reconstructed genome-scale metabolic model iSM454 for O. oeni PSU-1 and the resulting experimental data, we determined the redistribution of intracellular metabolic fluxes, under the different ethanol conditions. Four growth phases were clearly identified during the batch cultivation of O. oeni PSU-1 strain, according to the temporal consumption of malic and citric acids, sugar and amino acids uptake, and biosynthesis rates of metabolic products - biomass, erythritol, mannitol and acetic acid, among others. We showed that, under increasing ethanol conditions, O. oeni favors anabolic reactions related with cell maintenance, as the requirements of NAD(P)+ and ATP increased with ethanol content. Specifically, cultures containing 9 and 12% ethanol required 10 and 17 times more NGAM (non-growth associated maintenance ATP) during phase I, respectively, than cultures without ethanol. MLF and citric acid consumption are vital at high ethanol concentrations, as they are the main source for proton extrusion, allowing higher ATP production by F0F1-ATPase, the main route of ATP synthesis under these conditions. Mannitol and erythritol synthesis are the main sources of NAD(P)+, countervailing for 51-57% of its usage, as predicted by the model. Finally, cysteine shows the fastest specific consumption rate among the amino acids, confirming its key role for bacterial survival under ethanol stress. As a whole, this study provides a global insight into how ethanol content exerts a differential physiological response in O. oeni PSU-1 strain. It will help to design better strategies of nutrient addition to achieve a successful MLF of wine.
Keywords: Oenococcus oeni; genome-scale metabolic model; lactic acid bacteria; malolactic fermentation; physiological ethanol response; wine-like defined culture medium.
Figures

Similar articles
-
Metabolic behavior for a mutant Oenococcus oeni strain with high resistance to ethanol to survive under oenological multi-stress conditions.Front Microbiol. 2023 Mar 9;14:1100501. doi: 10.3389/fmicb.2023.1100501. eCollection 2023. Front Microbiol. 2023. PMID: 36970676 Free PMC article.
-
Genome-Scale Reconstruction of the Metabolic Network in Oenococcus oeni to Assess Wine Malolactic Fermentation.Front Microbiol. 2017 Mar 30;8:534. doi: 10.3389/fmicb.2017.00534. eCollection 2017. Front Microbiol. 2017. PMID: 28424673 Free PMC article.
-
Analysis of Transcriptomic Response to SO2 by Oenococcus oeni Growing in Continuous Culture.Microbiol Spectr. 2021 Oct 31;9(2):e0115421. doi: 10.1128/Spectrum.01154-21. Epub 2021 Oct 6. Microbiol Spectr. 2021. PMID: 34612664 Free PMC article.
-
Improving Oenococcus oeni to overcome challenges of wine malolactic fermentation.Trends Biotechnol. 2015 Sep;33(9):547-53. doi: 10.1016/j.tibtech.2015.06.008. Epub 2015 Jul 18. Trends Biotechnol. 2015. PMID: 26197706 Review.
-
Genomic analysis of Oenococcus oeni PSU-1 and its relevance to winemaking.FEMS Microbiol Rev. 2005 Aug;29(3):465-75. doi: 10.1016/j.femsre.2005.04.011. FEMS Microbiol Rev. 2005. PMID: 16125008 Review.
Cited by
-
Metabolic Modeling of Wine Fermentation at Genome Scale.Methods Mol Biol. 2022;2399:395-454. doi: 10.1007/978-1-0716-1831-8_16. Methods Mol Biol. 2022. PMID: 35604565
-
Citrate metabolism in lactic acid bacteria: is there a beneficial effect for Oenococcus oeni in wine?Front Microbiol. 2024 Jan 4;14:1283220. doi: 10.3389/fmicb.2023.1283220. eCollection 2023. Front Microbiol. 2024. PMID: 38249489 Free PMC article. Review.
-
Oenococcus oeni Exopolysaccharide Biosynthesis, a Tool to Improve Malolactic Starter Performance.Front Microbiol. 2018 Jun 12;9:1276. doi: 10.3389/fmicb.2018.01276. eCollection 2018. Front Microbiol. 2018. PMID: 29946314 Free PMC article.
-
Metabolic behavior for a mutant Oenococcus oeni strain with high resistance to ethanol to survive under oenological multi-stress conditions.Front Microbiol. 2023 Mar 9;14:1100501. doi: 10.3389/fmicb.2023.1100501. eCollection 2023. Front Microbiol. 2023. PMID: 36970676 Free PMC article.
-
Quantitative proteomic analysis reveals the ethanologenic metabolism regulation of Ethanoligenens harbinense by exogenous ethanol addition.Biotechnol Biofuels. 2019 Jun 28;12:166. doi: 10.1186/s13068-019-1511-y. eCollection 2019. Biotechnol Biofuels. 2019. PMID: 31297154 Free PMC article.
References
-
- Bartowsky E. J. (2005). Oenococcus oeni and malolactic fermentation–moving into the molecular arena. Aust. J. Grape Wine Res. 11 174–187. 10.1111/j.1755-0238.2005.tb00286.x - DOI
-
- Bauer R., Dicks L. (2004). Control of malolactic fermentation in wine. A review. South Afr. J. Enol. Vitic. 25 74–88.
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

