Microbial genome analysis: the COG approach
- PMID: 28968633
- PMCID: PMC6781585
- DOI: 10.1093/bib/bbx117
Microbial genome analysis: the COG approach
Abstract
For the past 20 years, the Clusters of Orthologous Genes (COG) database had been a popular tool for microbial genome annotation and comparative genomics. Initially created for the purpose of evolutionary classification of protein families, the COG have been used, apart from straightforward functional annotation of sequenced genomes, for such tasks as (i) unification of genome annotation in groups of related organisms; (ii) identification of missing and/or undetected genes in complete microbial genomes; (iii) analysis of genomic neighborhoods, in many cases allowing prediction of novel functional systems; (iv) analysis of metabolic pathways and prediction of alternative forms of enzymes; (v) comparison of organisms by COG functional categories; and (vi) prioritization of targets for structural and functional characterization. Here we review the principles of the COG approach and discuss its key advantages and drawbacks in microbial genome analysis.
Keywords: comparative genomics; enzyme evolution; genome annotation; orthologs; paralogs.
Published by Oxford University Press 2017. This work is written by US Government employees and is in the public domain in the US.
Figures

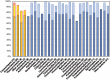
Similar articles
-
Expanded microbial genome coverage and improved protein family annotation in the COG database.Nucleic Acids Res. 2015 Jan;43(Database issue):D261-9. doi: 10.1093/nar/gku1223. Epub 2014 Nov 26. Nucleic Acids Res. 2015. PMID: 25428365 Free PMC article.
-
MicroScope-an integrated resource for community expertise of gene functions and comparative analysis of microbial genomic and metabolic data.Brief Bioinform. 2019 Jul 19;20(4):1071-1084. doi: 10.1093/bib/bbx113. Brief Bioinform. 2019. PMID: 28968784 Free PMC article.
-
Quantitatively Partitioning Microbial Genomic Traits among Taxonomic Ranks across the Microbial Tree of Life.mSphere. 2019 Aug 28;4(4):e00446-19. doi: 10.1128/mSphere.00446-19. mSphere. 2019. PMID: 31462411 Free PMC article.
-
Functional genomics and enzyme evolution. Homologous and analogous enzymes encoded in microbial genomes.Genetica. 1999;106(1-2):159-70. doi: 10.1023/a:1003705601428. Genetica. 1999. PMID: 10710722 Review.
-
Orthologs, paralogs, and evolutionary genomics.Annu Rev Genet. 2005;39:309-38. doi: 10.1146/annurev.genet.39.073003.114725. Annu Rev Genet. 2005. PMID: 16285863 Review.
Cited by
-
Microbiome and plant cell transformation trigger insect gall induction in cassava.Front Plant Sci. 2023 Nov 29;14:1237966. doi: 10.3389/fpls.2023.1237966. eCollection 2023. Front Plant Sci. 2023. PMID: 38126017 Free PMC article.
-
AYbRAH: a curated ortholog database for yeasts and fungi spanning 600 million years of evolution.Database (Oxford). 2019 Jan 1;2019:baz022. doi: 10.1093/database/baz022. Database (Oxford). 2019. PMID: 30893420 Free PMC article.
-
Dichloromethane Degradation Pathway from Unsequenced Hyphomicrobium sp. MC8b Rapidly Explored by Pan-Proteomics.Microorganisms. 2020 Nov 27;8(12):1876. doi: 10.3390/microorganisms8121876. Microorganisms. 2020. PMID: 33260855 Free PMC article.
-
How Rhizobia Adapt to the Nodule Environment.J Bacteriol. 2021 May 20;203(12):e0053920. doi: 10.1128/JB.00539-20. Epub 2021 Feb 1. J Bacteriol. 2021. PMID: 33526611 Free PMC article.
-
Estimating maximal microbial growth rates from cultures, metagenomes, and single cells via codon usage patterns.Proc Natl Acad Sci U S A. 2021 Mar 23;118(12):e2016810118. doi: 10.1073/pnas.2016810118. Proc Natl Acad Sci U S A. 2021. PMID: 33723043 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

