MiSynPat: An integrated knowledge base linking clinical, genetic, and structural data for disease-causing mutations in human mitochondrial aminoacyl-tRNA synthetases
- PMID: 28608363
- PMCID: PMC5638098
- DOI: 10.1002/humu.23277
MiSynPat: An integrated knowledge base linking clinical, genetic, and structural data for disease-causing mutations in human mitochondrial aminoacyl-tRNA synthetases
Abstract
Numerous mutations in each of the mitochondrial aminoacyl-tRNA synthetases (aaRSs) have been implicated in human diseases. The mutations are autosomal and recessive and lead mainly to neurological disorders, although with pleiotropic effects. The processes and interactions that drive the etiology of the disorders associated with mitochondrial aaRSs (mt-aaRSs) are far from understood. The complexity of the clinical, genetic, and structural data requires concerted, interdisciplinary efforts to understand the molecular biology of these disorders. Toward this goal, we designed MiSynPat, a comprehensive knowledge base together with an ergonomic Web server designed to organize and access all pertinent information (sequences, multiple sequence alignments, structures, disease descriptions, mutation characteristics, original literature) on the disease-linked human mt-aaRSs. With MiSynPat, a user can also evaluate the impact of a possible mutation on sequence-conservation-structure in order to foster the links between basic and clinical researchers and to facilitate future diagnosis. The proposed integrated view, coupled with research on disease-related mt-aaRSs, will help to reveal new functions for these enzymes and to open new vistas in the molecular biology of the cell. The purpose of MiSynPat, freely available at http://misynpat.org, is to constitute a reference and a converging resource for scientists and clinicians.
Keywords: 3D structures; aminoacyl-tRNA synthetases; disease-causing mutations; knowledge base; mitochondrial disorders; sequence alignments.
© 2017 The Authors. Human Mutation published by Wiley Periodicals, Inc.
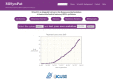
Figures

Similar articles
-
Evolutionary and structural annotation of disease-associated mutations in human aminoacyl-tRNA synthetases.BMC Genomics. 2014 Dec 4;15(1):1063. doi: 10.1186/1471-2164-15-1063. BMC Genomics. 2014. PMID: 25476837 Free PMC article.
-
Pathogenic implications of human mitochondrial aminoacyl-tRNA synthetases.Top Curr Chem. 2014;344:247-92. doi: 10.1007/128_2013_457. Top Curr Chem. 2014. PMID: 23824528 Review.
-
When a common biological role does not imply common disease outcomes: Disparate pathology linked to human mitochondrial aminoacyl-tRNA synthetases.J Biol Chem. 2019 Apr 5;294(14):5309-5320. doi: 10.1074/jbc.REV118.002953. Epub 2019 Jan 15. J Biol Chem. 2019. PMID: 30647134 Free PMC article. Review.
-
Intra-protein compensatory mutations analysis highlights the tRNA recognition regions in aminoacyl-tRNA synthetases.J Biomol Struct Dyn. 2009 Oct;27(2):115-26. doi: 10.1080/07391102.2009.10507302. J Biomol Struct Dyn. 2009. PMID: 19583438
-
Aminoacyl tRNA synthetases as malarial drug targets: a comparative bioinformatics study.Malar J. 2019 Feb 6;18(1):34. doi: 10.1186/s12936-019-2665-6. Malar J. 2019. PMID: 30728021 Free PMC article.
Cited by
-
Aminoacyl-tRNA synthetases in human health and disease.Front Physiol. 2022 Oct 18;13:1029218. doi: 10.3389/fphys.2022.1029218. eCollection 2022. Front Physiol. 2022. PMID: 36330207 Free PMC article. Review.
-
Clinical, neuroradiological, and molecular characterization of mitochondrial threonyl-tRNA-synthetase (TARS2)-related disorder.Genet Med. 2023 Nov;25(11):100938. doi: 10.1016/j.gim.2023.100938. Epub 2023 Jul 13. Genet Med. 2023. PMID: 37454282 Free PMC article.
-
Mitochondrial Aminoacyl-tRNA Synthetase and Disease: The Yeast Contribution for Functional Analysis of Novel Variants.Int J Mol Sci. 2021 Apr 26;22(9):4524. doi: 10.3390/ijms22094524. Int J Mol Sci. 2021. PMID: 33926074 Free PMC article. Review.
-
Mitochondrial aminoacyl-tRNA synthetase disorders: an emerging group of developmental disorders of myelination.J Neurodev Disord. 2019 Dec 16;11(1):29. doi: 10.1186/s11689-019-9292-y. J Neurodev Disord. 2019. PMID: 31839000 Free PMC article. Review.
-
Mitonuclear conflict and cooperation govern the integration of genotypes, phenotypes and environments.Philos Trans R Soc Lond B Biol Sci. 2020 Jan 20;375(1790):20190188. doi: 10.1098/rstb.2019.0188. Epub 2019 Dec 2. Philos Trans R Soc Lond B Biol Sci. 2020. PMID: 31787039 Free PMC article.
References
-
- Bonnefond, L. , Fender, A. , Rudinger‐Thirion, J. , Giegé, R. , Florentz, C. , & Sissler, M. (2005). Toward the full set of human mitochondrial aminoacyl‐tRNA synthetases: Characterization of AspRS and TyrRS. Biochemistry, 44, 4805–4816. - PubMed
-
- Bonnefond, L. , Frugier, M. , Touzé, E. , Lorber, B. , Florentz, C. , Giegé, R. , … Rudinger‐Thirion, J. (2007). Crystal structure of human mitochondrial tyrosyl‐tRNA synthetase reveals common and idiosyncratic features. Structure, 15(11)1505–1516. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

