Covalent targeting of remote cysteine residues to develop CDK12 and CDK13 inhibitors
- PMID: 27571479
- PMCID: PMC5033074
- DOI: 10.1038/nchembio.2166
Covalent targeting of remote cysteine residues to develop CDK12 and CDK13 inhibitors
Abstract
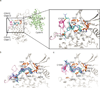
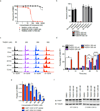
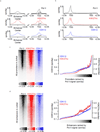
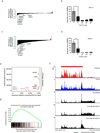
Cyclin-dependent kinases 12 and 13 (CDK12 and CDK13) play critical roles in the regulation of gene transcription. However, the absence of CDK12 and CDK13 inhibitors has hindered the ability to investigate the consequences of their inhibition in healthy cells and cancer cells. Here we describe the rational design of a first-in-class CDK12 and CDK13 covalent inhibitor, THZ531. Co-crystallization of THZ531 with CDK12-cyclin K indicates that THZ531 irreversibly targets a cysteine located outside the kinase domain. THZ531 causes a loss of gene expression with concurrent loss of elongating and hyperphosphorylated RNA polymerase II. In particular, THZ531 substantially decreases the expression of DNA damage response genes and key super-enhancer-associated transcription factor genes. Coincident with transcriptional perturbation, THZ531 dramatically induced apoptotic cell death. Small molecules capable of specifically targeting CDK12 and CDK13 may thus help identify cancer subtypes that are particularly dependent on their kinase activities.
Figures


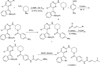

Comment in
-
Transcriptional kinases: Caught by a sticky drug.Nat Chem Biol. 2016 Sep 20;12(10):765-6. doi: 10.1038/nchembio.2184. Nat Chem Biol. 2016. PMID: 27649053 No abstract available.
Similar articles
-
Structure-activity relationship study of THZ531 derivatives enables the discovery of BSJ-01-175 as a dual CDK12/13 covalent inhibitor with efficacy in Ewing sarcoma.Eur J Med Chem. 2021 Oct 5;221:113481. doi: 10.1016/j.ejmech.2021.113481. Epub 2021 Apr 20. Eur J Med Chem. 2021. PMID: 33945934
-
Targeting CDK12-mediated transcription regulation in anaplastic thyroid carcinoma.Biochem Biophys Res Commun. 2019 Dec 10;520(3):544-550. doi: 10.1016/j.bbrc.2019.10.052. Epub 2019 Oct 12. Biochem Biophys Res Commun. 2019. PMID: 31615655
-
Noncovalent CDK12/13 dual inhibitors-based PROTACs degrade CDK12-Cyclin K complex and induce synthetic lethality with PARP inhibitor.Eur J Med Chem. 2022 Jan 15;228:114012. doi: 10.1016/j.ejmech.2021.114012. Epub 2021 Nov 25. Eur J Med Chem. 2022. PMID: 34864331
-
The promise and current status of CDK12/13 inhibition for the treatment of cancer.Future Med Chem. 2021 Jan;13(2):117-141. doi: 10.4155/fmc-2020-0240. Epub 2020 Dec 9. Future Med Chem. 2021. PMID: 33295810 Review.
-
Human CDK12 and CDK13, multi-tasking CTD kinases for the new millenium.Transcription. 2019 Apr;10(2):91-110. doi: 10.1080/21541264.2018.1535211. Epub 2018 Oct 22. Transcription. 2019. PMID: 30319007 Free PMC article. Review.
Cited by
-
Targeting the transcription cycle and RNA processing in cancer treatment.Curr Opin Pharmacol. 2021 Jun;58:69-75. doi: 10.1016/j.coph.2021.04.001. Epub 2021 May 5. Curr Opin Pharmacol. 2021. PMID: 33964728 Free PMC article. Review.
-
Therapeutic targeting of CPSF3-dependent transcriptional termination in ovarian cancer.Sci Adv. 2023 Nov 24;9(47):eadj0123. doi: 10.1126/sciadv.adj0123. Epub 2023 Nov 22. Sci Adv. 2023. PMID: 37992178 Free PMC article.
-
Inflammatory Immune-Associated eRNA: Mechanisms, Functions and Therapeutic Prospects.Front Immunol. 2022 Apr 19;13:849451. doi: 10.3389/fimmu.2022.849451. eCollection 2022. Front Immunol. 2022. PMID: 35514959 Free PMC article. Review.
-
PROTACs: great opportunities for academia and industry (an update from 2020 to 2021).Signal Transduct Target Ther. 2022 Jun 9;7(1):181. doi: 10.1038/s41392-022-00999-9. Signal Transduct Target Ther. 2022. PMID: 35680848 Free PMC article. Review.
-
Leveraging Compound Promiscuity to Identify Targetable Cysteines within the Kinome.Cell Chem Biol. 2019 Jun 20;26(6):818-829.e9. doi: 10.1016/j.chembiol.2019.02.021. Epub 2019 Apr 11. Cell Chem Biol. 2019. PMID: 30982749 Free PMC article.
References
Methods-only References
-
- Czudnochowski N, Bosken CA, Geyer M. Serine-7 but not serine-5 phosphorylation primes RNA polymerase II CTD for P-TEFb recognition. Nature communications. 2012;3:842. - PubMed
-
- Zhang Z, Marshall AGA. universal algorithm for fast and automated charge state deconvolution of electrospray mass-to-charge ratio spectra. Journal of the American Society for Mass Spectrometry. 1998;9:225–233. - PubMed
Publication types
MeSH terms
Substances
Associated data
- PubChem-Substance/316891618
- PubChem-Substance/316891619
- PubChem-Substance/316891620
- PubChem-Substance/316891621
- PubChem-Substance/316891622
- PubChem-Substance/316891623
- PubChem-Substance/316891624
- PubChem-Substance/316891625
- PubChem-Substance/316891626
- PubChem-Substance/316891627
- PubChem-Substance/316891628
- PubChem-Substance/316891629
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

