A Minimal Threshold of c-di-GMP Is Essential for Fruiting Body Formation and Sporulation in Myxococcus xanthus
- PMID: 27214040
- PMCID: PMC4877007
- DOI: 10.1371/journal.pgen.1006080
A Minimal Threshold of c-di-GMP Is Essential for Fruiting Body Formation and Sporulation in Myxococcus xanthus
Abstract
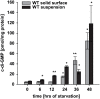
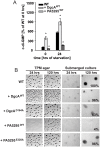
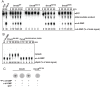
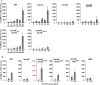
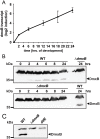
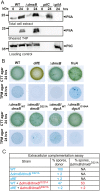
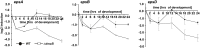
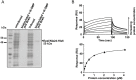
Generally, the second messenger bis-(3'-5')-cyclic dimeric GMP (c-di-GMP) regulates the switch between motile and sessile lifestyles in bacteria. Here, we show that c-di-GMP is an essential regulator of multicellular development in the social bacterium Myxococcus xanthus. In response to starvation, M. xanthus initiates a developmental program that culminates in formation of spore-filled fruiting bodies. We show that c-di-GMP accumulates at elevated levels during development and that this increase is essential for completion of development whereas excess c-di-GMP does not interfere with development. MXAN3735 (renamed DmxB) is identified as a diguanylate cyclase that only functions during development and is responsible for this increased c-di-GMP accumulation. DmxB synthesis is induced in response to starvation, thereby restricting DmxB activity to development. DmxB is essential for development and functions downstream of the Dif chemosensory system to stimulate exopolysaccharide accumulation by inducing transcription of a subset of the genes encoding proteins involved in exopolysaccharide synthesis. The developmental defects in the dmxB mutant are non-cell autonomous and rescued by co-development with a strain proficient in exopolysaccharide synthesis, suggesting reduced exopolysaccharide accumulation as the causative defect in this mutant. The NtrC-like transcriptional regulator EpsI/Nla24, which is required for exopolysaccharide accumulation, is identified as a c-di-GMP receptor, and thus a putative target for DmxB generated c-di-GMP. Because DmxB can be-at least partially-functionally replaced by a heterologous diguanylate cyclase, these results altogether suggest a model in which a minimum threshold level of c-di-GMP is essential for the successful completion of multicellular development in M. xanthus.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures

Similar articles
-
CRP-Like Transcriptional Regulator MrpC Curbs c-di-GMP and 3',3'-cGAMP Nucleotide Levels during Development in Myxococcus xanthus.mBio. 2021 Feb 22;13(1):e0004422. doi: 10.1128/mbio.00044-22. Epub 2022 Feb 15. mBio. 2021. PMID: 35164555 Free PMC article.
-
Three PilZ Domain Proteins, PlpA, PixA, and PixB, Have Distinct Functions in Regulation of Motility and Development in Myxococcus xanthus.J Bacteriol. 2021 Jun 8;203(13):e0012621. doi: 10.1128/JB.00126-21. Epub 2021 Jun 8. J Bacteriol. 2021. PMID: 33875546 Free PMC article.
-
Cyclic Di-GMP Regulates Type IV Pilus-Dependent Motility in Myxococcus xanthus.J Bacteriol. 2015 Jun 29;198(1):77-90. doi: 10.1128/JB.00281-15. Print 2016 Jan 1. J Bacteriol. 2015. PMID: 26124238 Free PMC article.
-
Bacterial diguanylate cyclases: structure, function and mechanism in exopolysaccharide biofilm development.Biotechnol Adv. 2015 Jan-Feb;33(1):124-141. doi: 10.1016/j.biotechadv.2014.11.010. Epub 2014 Dec 10. Biotechnol Adv. 2015. PMID: 25499693 Review.
-
[Activity of cyclic diguanylate (c-di-GMP) in bacteria and the study of its derivatives].Yao Xue Xue Bao. 2012 Mar;47(3):307-12. Yao Xue Xue Bao. 2012. PMID: 22645753 Review. Chinese.
Cited by
-
CRP-Like Transcriptional Regulator MrpC Curbs c-di-GMP and 3',3'-cGAMP Nucleotide Levels during Development in Myxococcus xanthus.mBio. 2021 Feb 22;13(1):e0004422. doi: 10.1128/mbio.00044-22. Epub 2022 Feb 15. mBio. 2021. PMID: 35164555 Free PMC article.
-
Three PilZ Domain Proteins, PlpA, PixA, and PixB, Have Distinct Functions in Regulation of Motility and Development in Myxococcus xanthus.J Bacteriol. 2021 Jun 8;203(13):e0012621. doi: 10.1128/JB.00126-21. Epub 2021 Jun 8. J Bacteriol. 2021. PMID: 33875546 Free PMC article.
-
Myxococcus xanthus PilB interacts with c-di-GMP and modulates motility and biofilm formation.J Bacteriol. 2023 Sep 26;205(9):e0022123. doi: 10.1128/jb.00221-23. Epub 2023 Sep 11. J Bacteriol. 2023. PMID: 37695853 Free PMC article.
-
Polymertropism of rod-shaped bacteria: movement along aligned polysaccharide fibers.Sci Rep. 2017 Aug 11;7(1):7643. doi: 10.1038/s41598-017-07486-0. Sci Rep. 2017. PMID: 28801641 Free PMC article.
-
Sequence Conservation, Domain Architectures, and Phylogenetic Distribution of the HD-GYP Type c-di-GMP Phosphodiesterases.J Bacteriol. 2022 Apr 19;204(4):e0056121. doi: 10.1128/jb.00561-21. Epub 2021 Dec 20. J Bacteriol. 2022. PMID: 34928179 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

