Synergistic growth in bacteria depends on substrate complexity
- PMID: 26727898
- PMCID: PMC4822414
- DOI: 10.1007/s12275-016-5461-9
Synergistic growth in bacteria depends on substrate complexity
Abstract
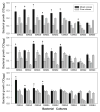
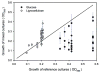
Both positive and negative interactions among bacteria take place in the environment. We hypothesize that the complexity of the substrate affects the way bacteria interact with greater cooperation in the presence of recalcitrant substrate. We isolated lignocellulolytic bacteria from salt marsh detritus and compared the growth, metabolic activity and enzyme production of pure cultures to those of three-species mixed cultures in lignocellulose and glucose media. Synergistic growth was common in lignocellulose medium containing carboxyl methyl cellulose, xylan and lignin but absent in glucose medium. Bacterial synergism promoted metabolic activity in synergistic mixed cultures but not the maximal growth rate (μ). Bacterial synergism also promoted the production of β-1,4-glucosidase but not the production of cellobiohydrolase or β-1,4-xylosidase. Our results suggest that the chemical complexity of the substrate affects the way bacteria interact. While a complex substrate such as lignocellulose promotes positive interactions and synergistic growth, a labile substrate such as glucose promotes negative interactions and competition. Synergistic interactions among indigenous bacteria are suggested to be important in promoting lignocellulose degradation in the environment.
Keywords: bacterial activity; bacterial synergism; enzyme production; lignocellulose degradation; microbial interaction.
Figures

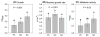
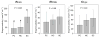
Similar articles
-
Influences of Media Compositions on Characteristics of Isolated Bacteria Exhibiting Lignocellulolytic Activities from Various Environmental Sites.Appl Biochem Biotechnol. 2017 Nov;183(3):931-942. doi: 10.1007/s12010-017-2474-8. Epub 2017 Apr 13. Appl Biochem Biotechnol. 2017. PMID: 28405916
-
Design and composition of synthetic fungal-bacterial microbial consortia that improve lignocellulolytic enzyme activity.Bioresour Technol. 2017 Mar;227:247-255. doi: 10.1016/j.biortech.2016.12.058. Epub 2016 Dec 22. Bioresour Technol. 2017. PMID: 28039824
-
Halotolerant microbial consortia able to degrade highly recalcitrant plant biomass substrate.Appl Microbiol Biotechnol. 2018 Mar;102(6):2913-2927. doi: 10.1007/s00253-017-8714-6. Epub 2018 Feb 3. Appl Microbiol Biotechnol. 2018. PMID: 29397428 Free PMC article.
-
[Research progress on plate mixed culture of lignocellulolytic microorganisms].Ying Yong Sheng Tai Xue Bao. 2012 Jun;23(6):1721-7. Ying Yong Sheng Tai Xue Bao. 2012. PMID: 22937666 Review. Chinese.
-
[Microbial degradation of lignocellulose].Sheng Wu Gong Cheng Xue Bao. 2019 Nov 25;35(11):2081-2091. doi: 10.13345/j.cjb.190248. Sheng Wu Gong Cheng Xue Bao. 2019. PMID: 31814356 Review. Chinese.
Cited by
-
Elucidating the Role of Biofilm-Forming Microbial Communities in Fermentative Biohydrogen Process: An Overview.Microorganisms. 2022 Sep 28;10(10):1924. doi: 10.3390/microorganisms10101924. Microorganisms. 2022. PMID: 36296200 Free PMC article. Review.
-
Bacterial Diversity and Community Structure of a Municipal Solid Waste Landfill: A Source of Lignocellulolytic Potential.Life (Basel). 2021 May 28;11(6):493. doi: 10.3390/life11060493. Life (Basel). 2021. PMID: 34071172 Free PMC article.
-
Long-Term Cellulose Enrichment Selects for Highly Cellulolytic Consortia and Competition for Public Goods.mSystems. 2022 Apr 26;7(2):e0151921. doi: 10.1128/msystems.01519-21. Epub 2022 Mar 8. mSystems. 2022. PMID: 35258341 Free PMC article.
-
Dynamics of carbon substrate competition among heterotrophic microorganisms.ISME J. 2024 Jan 8;18(1):wrae018. doi: 10.1093/ismejo/wrae018. ISME J. 2024. PMID: 38366177 Free PMC article.
-
A Review on Bacterial Contribution to Lignocellulose Breakdown into Useful Bio-Products.Int J Environ Res Public Health. 2021 Jun 3;18(11):6001. doi: 10.3390/ijerph18116001. Int J Environ Res Public Health. 2021. PMID: 34204975 Free PMC article. Review.
References
-
- Breugelmans P, Barken KB, Tolker-Nielsen T, Hofkens J, Dejonghe W, Springael D. Architecture and spatial organization in a triple-species bacterial biofilm synergistically degrading the phenylurea herbicide linuron. FEMS Microbiol Ecol. 2008;64:271–282. - PubMed
-
- Ciric L, Philp JC, Whiteley AS. Hydrocarbon utilization within a diesel-degrading bacterial consortium. FEMS Microbiol Lett. 2010;303:116–122. - PubMed
-
- D’Costa VM, Griffiths E, Wright GD. Expanding the soil antibiotic resistome: exploring environmental diversity. Curr Opin Microbiol. 2007;10:481–489. - PubMed
-
- Da Silva WJ, Seneviratne J, Parahitiyawa N, Rosa EA, Samaranayake LP, Del Bel Cury AA. Improvement of XTT assay performance for studies involving Candida albicans biofilms. Braz Dent J. 2008;19:364–369. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

