Review: Insights into molecular mechanisms of disease in neurodegeneration with brain iron accumulation: unifying theories
- PMID: 25870938
- PMCID: PMC4832581
- DOI: 10.1111/nan.12242
Review: Insights into molecular mechanisms of disease in neurodegeneration with brain iron accumulation: unifying theories
Abstract
Neurodegeneration with brain iron accumulation (NBIA) is a group of disorders characterized by dystonia, parkinsonism and spasticity. Iron accumulates in the basal ganglia and may be accompanied by Lewy bodies, axonal swellings and hyperphosphorylated tau depending on NBIA subtype. Mutations in 10 genes have been associated with NBIA that include Ceruloplasmin (Cp) and ferritin light chain (FTL), both directly involved in iron homeostasis, as well as Pantothenate Kinase 2 (PANK2), Phospholipase A2 group 6 (PLA2G6), Fatty acid hydroxylase 2 (FA2H), Coenzyme A synthase (COASY), C19orf12, WDR45 and DCAF17 (C2orf37). These genes are involved in seemingly unrelated cellular pathways, such as lipid metabolism, Coenzyme A synthesis and autophagy. A greater understanding of the cellular pathways that link these genes and the disease mechanisms leading to iron dyshomeostasis is needed. Additionally, the major overlap seen between NBIA and more common neurodegenerative diseases may highlight conserved disease processes. In this review, we will discuss clinical and pathological findings for each NBIA-related gene, discuss proposed disease mechanisms such as mitochondrial health, oxidative damage, autophagy/mitophagy and iron homeostasis, and speculate the potential overlap between NBIA subtypes.
Keywords: NBIA; Tau; autophagy; mitochondria; neurodegeneration; α-synuclein.
© 2015 The Authors. Neuropathology and Applied Neurobiology published by John Wiley & Sons Ltd on behalf of British Neuropathological Society.
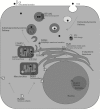
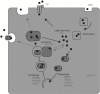
Figures
Similar articles
-
Neurodegeneration with brain iron accumulation: Insights into the mitochondria dysregulation.Biomed Pharmacother. 2019 Oct;118:109068. doi: 10.1016/j.biopha.2019.109068. Epub 2019 Aug 9. Biomed Pharmacother. 2019. PMID: 31404774 Review.
-
On the complexity of clinical and molecular bases of neurodegeneration with brain iron accumulation.Clin Genet. 2018 Apr;93(4):731-740. doi: 10.1111/cge.13057. Epub 2017 Sep 25. Clin Genet. 2018. PMID: 28542792 Review.
-
Mitochondrial Dysfunction, Oxidative Stress and Neuroinflammation in Neurodegeneration with Brain Iron Accumulation (NBIA).Antioxidants (Basel). 2020 Oct 20;9(10):1020. doi: 10.3390/antiox9101020. Antioxidants (Basel). 2020. PMID: 33092153 Free PMC article. Review.
-
Excess iron harms the brain: the syndromes of neurodegeneration with brain iron accumulation (NBIA).J Neural Transm (Vienna). 2013 Apr;120(4):695-703. doi: 10.1007/s00702-012-0922-8. Epub 2012 Dec 2. J Neural Transm (Vienna). 2013. PMID: 23212724 Review.
-
Neurodegeneration with brain iron accumulation.Handb Clin Neurol. 2018;147:293-305. doi: 10.1016/B978-0-444-63233-3.00019-1. Handb Clin Neurol. 2018. PMID: 29325618 Free PMC article. Review.
Cited by
-
Iron Overload in Brain: Transport Mismatches, Microbleeding Events, and How Nanochelating Therapies May Counteract Their Effects.Int J Mol Sci. 2024 Feb 16;25(4):2337. doi: 10.3390/ijms25042337. Int J Mol Sci. 2024. PMID: 38397013 Free PMC article. Review.
-
Unraveling Novel Mechanisms of Neurodegeneration Through a Large-Scale Forward Genetic Screen in Drosophila.Front Genet. 2019 Jan 14;9:700. doi: 10.3389/fgene.2018.00700. eCollection 2018. Front Genet. 2019. PMID: 30693015 Free PMC article. Review.
-
Does Ceruloplasmin Defend Against Neurodegenerative Diseases?Curr Neuropharmacol. 2019;17(6):539-549. doi: 10.2174/1570159X16666180508113025. Curr Neuropharmacol. 2019. PMID: 29737252 Free PMC article. Review.
-
Disorders of metal metabolism.Transl Sci Rare Dis. 2017 Dec 18;2(3-4):101-139. doi: 10.3233/TRD-170015. Transl Sci Rare Dis. 2017. PMID: 29354481 Free PMC article. Review.
-
Early-onset progressive spastic paraplegia caused by a novel TUBB4A mutation: brain MRI and FDG-PET findings.J Neurol. 2016 Mar;263(3):591-3. doi: 10.1007/s00415-016-8020-8. Epub 2016 Jan 25. J Neurol. 2016. PMID: 26810722 No abstract available.
References
-
- Hill JM, Switzer RC. The regional distribution and cellular localization of iron in the rat brain. Neuroscience 1984; 11: 595–603 - PubMed
-
- Hallgren B, Sourander P. The effect of age on the non‐haemin iron in the human brain. J Neurochem 1958; 3: 41–51 - PubMed
-
- Crichton RR, Wilmet S, Legssyer R, Ward RJ. Molecular and cellular mechanisms of iron homeostasis and toxicity in mammalian cells. J Inorg Biochem 2002; 91: 9–18 - PubMed
-
- Rouault TA. Iron metabolism in the CNS: implications for neurodegenerative diseases. Nat Rev Neurosci Nature Publishing Group 2013; 14: 551–564. - PubMed
-
- Andrews NC, Schmidt PJ. Iron homeostasis. Annu Rev Physiol Ann Rev 2007; 69: 69–85 - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Miscellaneous

