Structure of the catalytic domain of EZH2 reveals conformational plasticity in cofactor and substrate binding sites and explains oncogenic mutations
- PMID: 24367611
- PMCID: PMC3868588
- DOI: 10.1371/journal.pone.0083737
Structure of the catalytic domain of EZH2 reveals conformational plasticity in cofactor and substrate binding sites and explains oncogenic mutations
Abstract
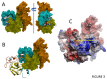
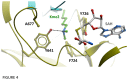
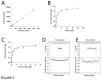
Polycomb repressive complex 2 (PRC2) is an important regulator of cellular differentiation and cell type identity. Overexpression or activating mutations of EZH2, the catalytic component of the PRC2 complex, are linked to hyper-trimethylation of lysine 27 of histone H3 (H3K27me3) in many cancers. Potent EZH2 inhibitors that reduce levels of H3K27me3 kill mutant lymphoma cells and are efficacious in a mouse xenograft model of malignant rhabdoid tumors. Unlike most SET domain methyltransferases, EZH2 requires PRC2 components, SUZ12 and EED, for activity, but the mechanism by which catalysis is promoted in the PRC2 complex is unknown. We solved the 2.0 Å crystal structure of the EZH2 methyltransferase domain revealing that most of the canonical structural features of SET domain methyltransferase structures are conserved. The site of methyl transfer is in a catalytically competent state, and the structure clarifies the structural mechanism underlying oncogenic hyper-trimethylation of H3K27 in tumors harboring mutations at Y641 or A677. On the other hand, the I-SET and post-SET domains occupy atypical positions relative to the core SET domain resulting in incomplete formation of the cofactor binding site and occlusion of the substrate binding groove. A novel CXC domain N-terminal to the SET domain may contribute to the apparent inactive conformation. We propose that protein interactions within the PRC2 complex modulate the trajectory of the post-SET and I-SET domains of EZH2 in favor of a catalytically competent conformation.
Conflict of interest statement
Figures


Similar articles
-
Structural context of disease-associated mutations and putative mechanism of autoinhibition revealed by X-ray crystallographic analysis of the EZH2-SET domain.PLoS One. 2013 Dec 19;8(12):e84147. doi: 10.1371/journal.pone.0084147. eCollection 2013. PLoS One. 2013. PMID: 24367637 Free PMC article.
-
Structural basis of oncogenic histone H3K27M inhibition of human polycomb repressive complex 2.Nat Commun. 2016 Apr 28;7:11316. doi: 10.1038/ncomms11316. Nat Commun. 2016. PMID: 27121947 Free PMC article.
-
EZH2 W113C is a gain-of-function mutation in B-cell lymphoma enabling both PRC2 methyltransferase activation and tazemetostat resistance.J Biol Chem. 2023 Apr;299(4):103073. doi: 10.1016/j.jbc.2023.103073. Epub 2023 Feb 27. J Biol Chem. 2023. PMID: 36858198 Free PMC article.
-
Structure, mechanism, and regulation of polycomb-repressive complex 2.J Biol Chem. 2018 Sep 7;293(36):13805-13814. doi: 10.1074/jbc.R117.800367. Epub 2017 Sep 14. J Biol Chem. 2018. PMID: 28912274 Free PMC article. Review.
-
Targeting EZH2 and PRC2 dependence as novel anticancer therapy.Exp Hematol. 2015 Aug;43(8):698-712. doi: 10.1016/j.exphem.2015.05.001. Epub 2015 May 28. Exp Hematol. 2015. PMID: 26027790 Free PMC article. Review.
Cited by
-
Molecular basis for chromatin assembly and modification by multiprotein complexes.Protein Sci. 2019 Feb;28(2):329-343. doi: 10.1002/pro.3535. Epub 2018 Dec 13. Protein Sci. 2019. PMID: 30350439 Free PMC article. Review.
-
Roles of H3K36-specific histone methyltransferases in transcription: antagonizing silencing and safeguarding transcription fidelity.Biophys Rep. 2018;4(4):170-177. doi: 10.1007/s41048-018-0063-1. Epub 2018 Aug 29. Biophys Rep. 2018. PMID: 30310854 Free PMC article. Review.
-
Human SETMAR is a DNA sequence-specific histone-methylase with a broad effect on the transcriptome.Nucleic Acids Res. 2019 Jan 10;47(1):122-133. doi: 10.1093/nar/gky937. Nucleic Acids Res. 2019. PMID: 30329085 Free PMC article.
-
Polycomb repressive complex 2 structure with inhibitor reveals a mechanism of activation and drug resistance.Nat Commun. 2016 Apr 28;7:11384. doi: 10.1038/ncomms11384. Nat Commun. 2016. PMID: 27122193 Free PMC article.
-
Maintaining cell identity: PRC2-mediated regulation of transcription and cancer.Nat Rev Cancer. 2016 Dec;16(12):803-810. doi: 10.1038/nrc.2016.83. Epub 2016 Sep 23. Nat Rev Cancer. 2016. PMID: 27658528 Review.
References
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

