Interaction between AP-5 and the hereditary spastic paraplegia proteins SPG11 and SPG15
- PMID: 23825025
- PMCID: PMC3744948
- DOI: 10.1091/mbc.E13-03-0170
Interaction between AP-5 and the hereditary spastic paraplegia proteins SPG11 and SPG15
Abstract
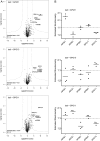
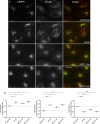
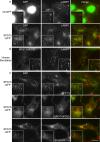
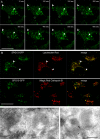
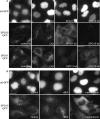
The AP-5 complex is a recently identified but evolutionarily ancient member of the family of heterotetrameric adaptor proteins (AP complexes). It is associated with two proteins that are mutated in patients with hereditary spastic paraplegia, SPG11 and SPG15. Here we show that the four AP-5 subunits can be coimmunoprecipitated with SPG11 and SPG15, both from cytosol and from detergent-extracted membranes, with a stoichiometry of ∼1:1:1:1:1:1. Knockdowns of SPG11 or SPG15 phenocopy knockdowns of AP-5 subunits: all six knockdowns cause the cation-independent mannose 6-phosphate receptor to become trapped in clusters of early endosomes. In addition, AP-5, SPG11, and SPG15 colocalize on a late endosomal/lysosomal compartment. Both SPG11 and SPG15 have predicted secondary structures containing α-solenoids related to those of clathrin heavy chain and COPI subunits. SPG11 also has an N-terminal, β-propeller-like domain, which interacts in vitro with AP-5. We propose that AP-5, SPG15, and SPG11 form a coat-like complex, with AP-5 involved in protein sorting, SPG15 facilitating the docking of the coat onto membranes by interacting with PI3P via its FYVE domain, and SPG11 (possibly together with SPG15) forming a scaffold.
Figures


Similar articles
-
ZFYVE26/SPASTIZIN and SPG11/SPATACSIN mutations in hereditary spastic paraplegia types AR-SPG15 and AR-SPG11 have different effects on autophagy and endocytosis.Autophagy. 2019 Jan;15(1):34-57. doi: 10.1080/15548627.2018.1507438. Epub 2018 Sep 13. Autophagy. 2019. PMID: 30081747 Free PMC article.
-
Rag GTPases and phosphatidylinositol 3-phosphate mediate recruitment of the AP-5/SPG11/SPG15 complex.J Cell Biol. 2021 Feb 1;220(2):e202002075. doi: 10.1083/jcb.202002075. J Cell Biol. 2021. PMID: 33464297 Free PMC article.
-
Frequency and phenotype of SPG11 and SPG15 in complicated hereditary spastic paraplegia.J Neurol Neurosurg Psychiatry. 2009 Dec;80(12):1402-4. doi: 10.1136/jnnp.2008.167528. J Neurol Neurosurg Psychiatry. 2009. PMID: 19917823
-
Adaptor protein complexes AP-4 and AP-5: new players in endosomal trafficking and progressive spastic paraplegia.Traffic. 2013 Feb;14(2):153-64. doi: 10.1111/tra.12028. Epub 2012 Dec 7. Traffic. 2013. PMID: 23167973 Review.
-
The role of AP-4 in cargo export from the trans-Golgi network and hereditary spastic paraplegia.Biochem Soc Trans. 2020 Oct 30;48(5):1877-1888. doi: 10.1042/BST20190664. Biochem Soc Trans. 2020. PMID: 33084855 Review.
Cited by
-
Role of the AP-5 adaptor protein complex in late endosome-to-Golgi retrieval.PLoS Biol. 2018 Jan 30;16(1):e2004411. doi: 10.1371/journal.pbio.2004411. eCollection 2018 Jan. PLoS Biol. 2018. PMID: 29381698 Free PMC article.
-
Souffle/Spastizin controls secretory vesicle maturation during zebrafish oogenesis.PLoS Genet. 2014 Jun 26;10(6):e1004449. doi: 10.1371/journal.pgen.1004449. eCollection 2014 Jun. PLoS Genet. 2014. PMID: 24967841 Free PMC article.
-
Neuronal lysosomes.Neurosci Lett. 2019 Apr 1;697:1-9. doi: 10.1016/j.neulet.2018.04.005. Epub 2018 Apr 4. Neurosci Lett. 2019. PMID: 29626653 Free PMC article. Review.
-
Altered Metabolism in Motor Neuron Diseases: Mechanism and Potential Therapeutic Target.Cells. 2023 Jun 2;12(11):1536. doi: 10.3390/cells12111536. Cells. 2023. PMID: 37296656 Free PMC article. Review.
-
AP-4 mediates export of ATG9A from the trans-Golgi network to promote autophagosome formation.Proc Natl Acad Sci U S A. 2017 Dec 12;114(50):E10697-E10706. doi: 10.1073/pnas.1717327114. Epub 2017 Nov 27. Proc Natl Acad Sci U S A. 2017. PMID: 29180427 Free PMC article.
References
-
- Arike L, Valgepea K, Peil L, Nahku R, Adamberg K, Vilu R. Comparison and applications of label-free absolute proteome quantification methods on Escherichia coli. J Proteomics. 2012;75:5437–5448. - PubMed
-
- Beck KA, Keen JH. Interaction of phosphoinositide cycle intermediates with the plasma membrane-associated clathrin assembly protein AP-2. J Biol Chem. 1991;266:4442–4447. - PubMed
-
- Bettencourt C, et al. Exome sequencing is a useful diagnostic tool for complicated forms of hereditary spastic paraplegia. Clin Genet. 2013 doi: 10.1111/cge.12133. - PubMed
-
- Boukhris A, et al. Hereditary spastic paraplegia with mental impairment and thin corpus callosum in Tunisia: SPG11, SPG15, and further genetic heterogeneity. Arch Neurol. 2008;65:393–402. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

