Specific Notch receptor-ligand interactions control human TCR-αβ/γδ development by inducing differential Notch signal strength
- PMID: 23530123
- PMCID: PMC3620353
- DOI: 10.1084/jem.20121798
Specific Notch receptor-ligand interactions control human TCR-αβ/γδ development by inducing differential Notch signal strength
Abstract
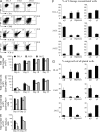
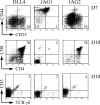
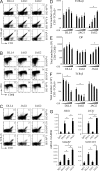
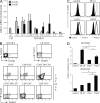
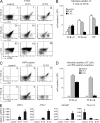
In humans, high Notch activation promotes γδ T cell development, whereas lower levels promote αβ-lineage differentiation. How these different Notch signals are generated has remained unclear. We show that differential Notch receptor-ligand interactions mediate this process. Whereas Delta-like 4 supports both TCR-αβ and -γδ development, Jagged1 induces mainly αβ-lineage differentiation. In contrast, Jagged2-mediated Notch activation primarily results in γδ T cell development and represses αβ-lineage differentiation by inhibiting TCR-β formation. Consistently, TCR-αβ T cell development is rescued through transduction of a TCR-β transgene. Jagged2 induces the strongest Notch signal through interactions with both Notch1 and Notch3, whereas Delta-like 4 primarily binds Notch1. In agreement, Notch3 is a stronger Notch activator and only supports γδ T cell development, whereas Notch1 is a weaker activator supporting both TCR-αβ and -γδ development. Fetal thymus organ cultures in JAG2-deficient thymic lobes or with Notch3-blocking antibodies confirm the importance of Jagged2/Notch3 signaling in human TCR-γδ differentiation. Our findings reveal that differential Notch receptor-ligand interactions mediate human TCR-αβ and -γδ T cell differentiation and provide a mechanistic insight into the high Notch dependency of human γδ T cell development.
Figures



Similar articles
-
Delta-like 4-mediated Notch signaling is required for early T-cell development in a three-dimensional thymic structure.Eur J Immunol. 2015 Aug;45(8):2252-62. doi: 10.1002/eji.201445123. Epub 2015 Jun 3. Eur J Immunol. 2015. PMID: 25976373
-
An early decrease in Notch activation is required for human TCR-alphabeta lineage differentiation at the expense of TCR-gammadelta T cells.Blood. 2009 Mar 26;113(13):2988-98. doi: 10.1182/blood-2008-06-164871. Epub 2008 Dec 3. Blood. 2009. PMID: 19056690
-
Expression pattern of notch1, 2 and 3 and Jagged1 and 2 in lymphoid and stromal thymus components: distinct ligand-receptor interactions in intrathymic T cell development.Int Immunol. 1999 Jul;11(7):1017-25. doi: 10.1093/intimm/11.7.1017. Int Immunol. 1999. PMID: 10383933
-
αβ versus γδ fate choice: counting the T-cell lineages at the branch point.Immunol Rev. 2010 Nov;238(1):169-81. doi: 10.1111/j.1600-065X.2010.00947.x. Immunol Rev. 2010. PMID: 20969592 Free PMC article. Review.
-
αβ and γδ T cell receptors: Similar but different.J Leukoc Biol. 2020 Jun;107(6):1045-1055. doi: 10.1002/JLB.2MR1219-233R. Epub 2020 Jan 29. J Leukoc Biol. 2020. PMID: 31994778 Review.
Cited by
-
Characterization of the genome-wide TLX1 binding profile in T-cell acute lymphoblastic leukemia.Leukemia. 2015 Dec;29(12):2317-27. doi: 10.1038/leu.2015.162. Epub 2015 Jun 25. Leukemia. 2015. PMID: 26108691
-
The Jekyll and Hyde story of IL17-Producing γδT Cells.Front Immunol. 2015 Feb 4;6:37. doi: 10.3389/fimmu.2015.00037. eCollection 2015. Front Immunol. 2015. PMID: 25699053 Free PMC article. Review.
-
Programming for T-lymphocyte fates: modularity and mechanisms.Genes Dev. 2019 Sep 1;33(17-18):1117-1135. doi: 10.1101/gad.327163.119. Genes Dev. 2019. PMID: 31481536 Free PMC article. Review.
-
[JAG1 promotes migration, invasion, and adhesion of triple-negative breast cancer cells by promoting angiogenesis].Nan Fang Yi Ke Da Xue Xue Bao. 2022 Jul 20;42(7):1100-1108. doi: 10.12122/j.issn.1673-4254.2022.07.21. Nan Fang Yi Ke Da Xue Xue Bao. 2022. PMID: 35869777 Free PMC article. Chinese.
-
NOTCH Signaling in Osteosarcoma.Curr Issues Mol Biol. 2023 Mar 8;45(3):2266-2283. doi: 10.3390/cimb45030146. Curr Issues Mol Biol. 2023. PMID: 36975516 Free PMC article. Review.
References
-
- Beatus P., Lundkvist J., Oberg C., Lendahl U. 1999. The notch 3 intracellular domain represses notch 1-mediated activation through Hairy/Enhancer of split (HES) promoters. Development. 126:3925–3935 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

