Derepression of a neuronal inhibitor due to miRNA dysregulation in a schizophrenia-related microdeletion
- PMID: 23332760
- PMCID: PMC3556818
- DOI: 10.1016/j.cell.2012.11.052
Derepression of a neuronal inhibitor due to miRNA dysregulation in a schizophrenia-related microdeletion
Abstract
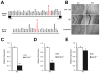
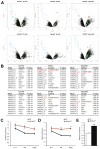
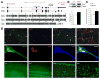
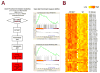
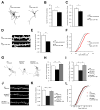
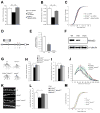
22q11.2 microdeletions result in specific cognitive deficits and schizophrenia. Analysis of Df(16)A(+/-) mice, which model this microdeletion, revealed abnormalities in the formation of neuronal dendrites and spines, as well as altered brain microRNAs. Here, we show a drastic reduction of miR-185, which resides within the 22q11.2 locus, to levels more than expected by a hemizygous deletion, and we demonstrate that this reduction alters dendritic and spine development. miR-185 represses, through an evolutionarily conserved target site, a previously unknown inhibitor of these processes that resides in the Golgi apparatus and shows higher prenatal brain expression. Sustained derepression of this inhibitor after birth represents the most robust transcriptional disturbance in the brains of Df(16)A(+/-) mice and results in structural alterations in the hippocampus. Reduction of miR-185 also has milder age- and region-specific effects on the expression of some Golgi-related genes. Our findings illuminate the contribution of microRNAs in psychiatric disorders and cognitive dysfunction.
Copyright © 2013 Elsevier Inc. All rights reserved.
Figures

Similar articles
-
Palmitoylation-dependent neurodevelopmental deficits in a mouse model of 22q11 microdeletion.Nat Neurosci. 2008 Nov;11(11):1302-10. doi: 10.1038/nn.2204. Epub 2008 Oct 5. Nat Neurosci. 2008. PMID: 18836441 Free PMC article.
-
Altered brain microRNA biogenesis contributes to phenotypic deficits in a 22q11-deletion mouse model.Nat Genet. 2008 Jun;40(6):751-60. doi: 10.1038/ng.138. Epub 2008 May 11. Nat Genet. 2008. PMID: 18469815
-
System-based proteomic and metabonomic analysis of the Df(16)A+/- mouse identifies potential miR-185 targets and molecular pathway alterations.Mol Psychiatry. 2017 Mar;22(3):384-395. doi: 10.1038/mp.2016.27. Epub 2016 Mar 22. Mol Psychiatry. 2017. PMID: 27001617 Free PMC article.
-
The 22q11.2 microdeletion: fifteen years of insights into the genetic and neural complexity of psychiatric disorders.Int J Dev Neurosci. 2011 May;29(3):259-81. doi: 10.1016/j.ijdevneu.2010.09.007. Epub 2010 Oct 8. Int J Dev Neurosci. 2011. PMID: 20920576 Free PMC article. Review.
-
22q11.2 microdeletions: linking DNA structural variation to brain dysfunction and schizophrenia.Nat Rev Neurosci. 2010 Jun;11(6):402-16. doi: 10.1038/nrn2841. Nat Rev Neurosci. 2010. PMID: 20485365 Free PMC article. Review.
Cited by
-
A Disc1 mutation differentially affects neurites and spines in hippocampal and cortical neurons.Mol Cell Neurosci. 2013 May;54:84-92. doi: 10.1016/j.mcn.2013.01.006. Epub 2013 Feb 6. Mol Cell Neurosci. 2013. PMID: 23396153 Free PMC article.
-
Temporal dynamics of miRNAs in human DLPFC and its association with miRNA dysregulation in schizophrenia.Transl Psychiatry. 2019 Aug 20;9(1):196. doi: 10.1038/s41398-019-0538-y. Transl Psychiatry. 2019. PMID: 31431609 Free PMC article.
-
Downregulation of miR-185 is a common pathogenic event in 22q11.2 deletion syndrome-related and idiopathic schizophrenia.Metab Brain Dis. 2022 Apr;37(4):1175-1184. doi: 10.1007/s11011-022-00918-5. Epub 2022 Jan 25. Metab Brain Dis. 2022. PMID: 35075501
-
Identifying lncRNA-miRNA-mRNA networks to investigate Alzheimer's disease pathogenesis and therapy strategy.Aging (Albany NY). 2020 Feb 7;12(3):2897-2920. doi: 10.18632/aging.102785. Epub 2020 Feb 7. Aging (Albany NY). 2020. PMID: 32035423 Free PMC article.
-
MicroRNAs: fundamental regulators of gene expression in major affective disorders and suicidal behavior?Front Cell Neurosci. 2013 Nov 15;7:208. doi: 10.3389/fncel.2013.00208. eCollection 2013. Front Cell Neurosci. 2013. PMID: 24298237 Free PMC article. No abstract available.
References
-
- Ambros V. MicroRNAs: genetically sensitized worms reveal new secrets. Curr Biol. 2010;20:R598–600. - PubMed
-
- Arguello PA, Gogos JA. Modeling madness in mice: one piece at a time. Nuron. 2006;52:179–196. - PubMed
-
- Arguello PA, Gogos JA. Genetic and cognitive windows into circuit mechanisms of psychiatric disease. Trends Neurosci. 2012;35:3–13. - PubMed
Publication types
MeSH terms
Substances
Supplementary concepts
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

