Exome capture reveals ZNF423 and CEP164 mutations, linking renal ciliopathies to DNA damage response signaling
- PMID: 22863007
- PMCID: PMC3433835
- DOI: 10.1016/j.cell.2012.06.028
Exome capture reveals ZNF423 and CEP164 mutations, linking renal ciliopathies to DNA damage response signaling
Abstract
Nephronophthisis-related ciliopathies (NPHP-RC) are degenerative recessive diseases that affect kidney, retina, and brain. Genetic defects in NPHP gene products that localize to cilia and centrosomes defined them as "ciliopathies." However, disease mechanisms remain poorly understood. Here, we identify by whole-exome resequencing, mutations of MRE11, ZNF423, and CEP164 as causing NPHP-RC. All three genes function within the DNA damage response (DDR) pathway. We demonstrate that, upon induced DNA damage, the NPHP-RC proteins ZNF423, CEP164, and NPHP10 colocalize to nuclear foci positive for TIP60, known to activate ATM at sites of DNA damage. We show that knockdown of CEP164 or ZNF423 causes sensitivity to DNA damaging agents and that cep164 knockdown in zebrafish results in dysregulated DDR and an NPHP-RC phenotype. Our findings link degenerative diseases of the kidney and retina, disorders of increasing prevalence, to mechanisms of DDR.
Copyright © 2012 Elsevier Inc. All rights reserved.
Conflict of interest statement
The authors declare that they have no competing financial interests.
Figures
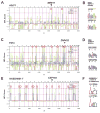
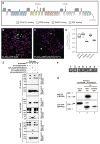
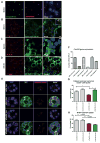

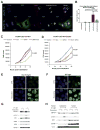
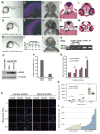
Similar articles
-
Nephronophthisis-associated CEP164 regulates cell cycle progression, apoptosis and epithelial-to-mesenchymal transition.PLoS Genet. 2014 Oct 23;10(10):e1004594. doi: 10.1371/journal.pgen.1004594. eCollection 2014 Oct. PLoS Genet. 2014. PMID: 25340510 Free PMC article.
-
Embryonic and foetal expression patterns of the ciliopathy gene CEP164.PLoS One. 2020 Jan 28;15(1):e0221914. doi: 10.1371/journal.pone.0221914. eCollection 2020. PLoS One. 2020. PMID: 31990917 Free PMC article.
-
Mutation of the Mg2+ transporter SLC41A1 results in a nephronophthisis-like phenotype.J Am Soc Nephrol. 2013 May;24(6):967-77. doi: 10.1681/ASN.2012101034. Epub 2013 May 9. J Am Soc Nephrol. 2013. PMID: 23661805 Free PMC article.
-
Nephronophthisis: a pathological and genetic perspective.Pediatr Nephrol. 2024 Jul;39(7):1977-2000. doi: 10.1007/s00467-023-06174-8. Epub 2023 Nov 6. Pediatr Nephrol. 2024. PMID: 37930417 Review.
-
Nephronophthisis-associated ciliopathies.J Am Soc Nephrol. 2007 Jun;18(6):1855-71. doi: 10.1681/ASN.2006121344. Epub 2007 May 18. J Am Soc Nephrol. 2007. PMID: 17513324 Review.
Cited by
-
Genetic Heterogeneity Underlying Phenotypes with Early-Onset Cerebellar Atrophy.Int J Mol Sci. 2023 Nov 16;24(22):16400. doi: 10.3390/ijms242216400. Int J Mol Sci. 2023. PMID: 38003592 Free PMC article.
-
Mutations of the SLIT2-ROBO2 pathway genes SLIT2 and SRGAP1 confer risk for congenital anomalies of the kidney and urinary tract.Hum Genet. 2015 Aug;134(8):905-16. doi: 10.1007/s00439-015-1570-5. Epub 2015 May 31. Hum Genet. 2015. PMID: 26026792 Free PMC article. Clinical Trial.
-
Nek1 kinase associates with ATR-ATRIP and primes ATR for efficient DNA damage signaling.Proc Natl Acad Sci U S A. 2013 Feb 5;110(6):2175-80. doi: 10.1073/pnas.1217781110. Epub 2013 Jan 23. Proc Natl Acad Sci U S A. 2013. PMID: 23345434 Free PMC article.
-
Deletion of CEP164 in mouse photoreceptors post-ciliogenesis interrupts ciliary intraflagellar transport (IFT).PLoS Genet. 2022 Sep 8;18(9):e1010154. doi: 10.1371/journal.pgen.1010154. eCollection 2022 Sep. PLoS Genet. 2022. PMID: 36074756 Free PMC article.
-
ZNF423: Transcriptional modulation in development and cancer.Mol Cell Oncol. 2014 Dec 23;1(3):e969655. doi: 10.4161/23723548.2014.969655. eCollection 2014 Jul-Sep. Mol Cell Oncol. 2014. PMID: 27308357 Free PMC article. Review.
References
-
- Andersen JS, Wilkinson CJ, Mayor T, Mortensen P, Nigg EA, Mann M. Proteomic characterization of the human centrosome by protein correlation profiling. Nature. 2003;426:570–574. - PubMed
-
- Angers S, Thorpe CJ, Biechele TL, Goldenberg SJ, Zheng N, MacCoss MJ, Moon RT. The KLHL12-Cullin-3 ubiquitin ligase negatively regulates the Wnt-beta-catenin pathway by targeting Dishevelled for degradation. Nat Cell Biol. 2006;8:348–357. - PubMed
-
- Ansley SJ, Badano JL, Blacque OE, Hill J, Hoskins BE, Leitch CC, Kim JC, Ross AJ, Eichers ER, Teslovich TM, et al. Basal body dysfunction is a likely cause of pleiotropic Bardet-Biedl syndrome. Nature. 2003;425:628–633. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- DK091405/DK/NIDDK NIH HHS/United States
- DK072301/DK/NIDDK NIH HHS/United States
- R01 DK088767/DK/NIDDK NIH HHS/United States
- R01 DK072301/DK/NIDDK NIH HHS/United States
- HHMI/Howard Hughes Medical Institute/United States
- S10 RR026550/RR/NCRR NIH HHS/United States
- R01 HL095545/HL/NHLBI NIH HHS/United States
- P60 DK020572/DK/NIDDK NIH HHS/United States
- R01 DK068306/DK/NIDDK NIH HHS/United States
- NS060109/NS/NINDS NIH HHS/United States
- P30 DK020572/DK/NIDDK NIH HHS/United States
- K99 DK091405/DK/NIDDK NIH HHS/United States
- DK068306/DK/NIDDK NIH HHS/United States
- F32 EY019430/EY/NEI NIH HHS/United States
- NS054871/NS/NINDS NIH HHS/United States
- F32EY19430/EY/NEI NIH HHS/United States
- G0700073/MRC_/Medical Research Council/United Kingdom
- R01 NS054871/NS/NINDS NIH HHS/United States
- R01EY018571/EY/NEI NIH HHS/United States
- U54 HG003273/HG/NHGRI NIH HHS/United States
- HD042601/HD/NICHD NIH HHS/United States
- R01 DK076683/DK/NIDDK NIH HHS/United States
- R01 DK075972/DK/NIDDK NIH HHS/United States
- DK090917/DK/NIDDK NIH HHS/United States
- DK075972/DK/NIDDK NIH HHS/United States
- R01 EY018571/EY/NEI NIH HHS/United States
- RC4 DK090917/DK/NIDDK NIH HHS/United States
- R01 HD042601/HD/NICHD NIH HHS/United States
- R01 NS060109/NS/NINDS NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

