Sensing the messenger: the diverse ways that bacteria signal through c-di-GMP
- PMID: 22593024
- PMCID: PMC3403432
- DOI: 10.1002/pro.2093
Sensing the messenger: the diverse ways that bacteria signal through c-di-GMP
Abstract
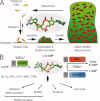
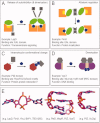
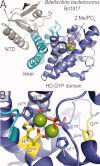
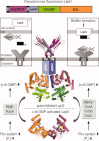
An intracellular second messenger unique to bacteria, c-di-GMP, has gained appreciation as a key player in adaptation and virulence strategies, such as biofilm formation, persistence, and cytotoxicity. Diguanylate cyclases containing GGDEF domains and phosphodiesterases containing either EAL or HD-GYP domains have been identified as the enzymes controlling intracellular c-di-GMP levels, yet little is known regarding signal transmission and the sensory targets for this signaling molecule. Although limited in number, identified c-di-GMP receptors in bacteria are characterized by prominent diversity and multilevel impact. In addition, c-di-GMP has been shown to have immunomodulatory effects in mammals and several eukaryotic c-di-GMP sensors have been proposed. The structural biology of c-di-GMP receptors is a rapidly developing field of research, which holds promise for the development of novel therapeutics against bacterial infections. In this review, we highlight recent advances in identifying bacterial and eukaryotic c-di-GMP signaling mechanisms and emphasize the need for mechanistic structure-function studies on confirmed signaling targets.
Copyright © 2012 The Protein Society.
Figures




Similar articles
-
In silico comparative analysis of GGDEF and EAL domain signaling proteins from the Azospirillum genomes.BMC Microbiol. 2018 Mar 9;18(1):20. doi: 10.1186/s12866-018-1157-0. BMC Microbiol. 2018. PMID: 29523074 Free PMC article.
-
A novel bacterial l-arginine sensor controlling c-di-GMP levels in Pseudomonas aeruginosa.Proteins. 2018 Oct;86(10):1088-1096. doi: 10.1002/prot.25587. Epub 2018 Sep 8. Proteins. 2018. PMID: 30040157
-
More than Enzymes That Make or Break Cyclic Di-GMP-Local Signaling in the Interactome of GGDEF/EAL Domain Proteins of Escherichia coli.mBio. 2017 Oct 10;8(5):e01639-17. doi: 10.1128/mBio.01639-17. mBio. 2017. PMID: 29018125 Free PMC article.
-
[Activity of cyclic diguanylate (c-di-GMP) in bacteria and the study of its derivatives].Yao Xue Xue Bao. 2012 Mar;47(3):307-12. Yao Xue Xue Bao. 2012. PMID: 22645753 Review. Chinese.
-
C-di-GMP Synthesis: Structural Aspects of Evolution, Catalysis and Regulation.J Mol Biol. 2016 Sep 25;428(19):3683-701. doi: 10.1016/j.jmb.2016.07.023. Epub 2016 Aug 4. J Mol Biol. 2016. PMID: 27498163 Review.
Cited by
-
Cyclic Di-GMP Regulates Type IV Pilus-Dependent Motility in Myxococcus xanthus.J Bacteriol. 2015 Jun 29;198(1):77-90. doi: 10.1128/JB.00281-15. Print 2016 Jan 1. J Bacteriol. 2015. PMID: 26124238 Free PMC article.
-
The Vibrio cholerae master regulator for the activation of biofilm biogenesis genes, VpsR, senses both cyclic di-GMP and phosphate.Nucleic Acids Res. 2022 May 6;50(8):4484-4499. doi: 10.1093/nar/gkac253. Nucleic Acids Res. 2022. PMID: 35438787 Free PMC article.
-
Oligoribonuclease is the primary degradative enzyme for pGpG in Pseudomonas aeruginosa that is required for cyclic-di-GMP turnover.Proc Natl Acad Sci U S A. 2015 Sep 8;112(36):E5048-57. doi: 10.1073/pnas.1507245112. Epub 2015 Aug 24. Proc Natl Acad Sci U S A. 2015. PMID: 26305945 Free PMC article.
-
The stringent response promotes biofilm dispersal in Pseudomonas putida.Sci Rep. 2017 Dec 22;7(1):18055. doi: 10.1038/s41598-017-18518-0. Sci Rep. 2017. PMID: 29273811 Free PMC article.
-
Local signaling enhances output specificity of bacterial c-di-GMP signaling networks.Microlife. 2023 May 11;4:uqad026. doi: 10.1093/femsml/uqad026. eCollection 2023. Microlife. 2023. PMID: 37251514 Free PMC article. Review.
References
-
- Ross P, Weinhouse H, Aloni Y, Michaeli D, Weinberger-Ohana P, Mayer R, Braun S, de Vroom E, van der Marel GA, van Boom JH, Benziman M. Regulation of cellulose synthesis in Acetobacter xylinum by cyclic diguanylic acid. Nature. 1987;325:279–281. - PubMed
-
- O'Toole G, Kaplan HB, Kolter R. Biofilm formation as microbial development. Annu Rev Microbiol. 2000;54:49–79. - PubMed
-
- Jenal U, Malone J. Mechanisms of cyclic-di-GMP signaling in bacteria. Annu Rev Genet. 2006;40:385–407. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

