Clinical application of exome sequencing in undiagnosed genetic conditions
- PMID: 22581936
- PMCID: PMC3375064
- DOI: 10.1136/jmedgenet-2012-100819
Clinical application of exome sequencing in undiagnosed genetic conditions
Abstract
Background: There is considerable interest in the use of next-generation sequencing to help diagnose unidentified genetic conditions, but it is difficult to predict the success rate in a clinical setting that includes patients with a broad range of phenotypic presentations.
Methods: The authors present a pilot programme of whole-exome sequencing on 12 patients with unexplained and apparent genetic conditions, along with their unaffected parents. Unlike many previous studies, the authors did not seek patients with similar phenotypes, but rather enrolled any undiagnosed proband with an apparent genetic condition when predetermined criteria were met.
Results: This undertaking resulted in a likely genetic diagnosis in 6 of the 12 probands, including the identification of apparently causal mutations in four genes known to cause Mendelian disease (TCF4, EFTUD2, SCN2A and SMAD4) and one gene related to known Mendelian disease genes (NGLY1). Of particular interest is that at the time of this study, EFTUD2 was not yet known as a Mendelian disease gene but was nominated as a likely cause based on the observation of de novo mutations in two unrelated probands. In a seventh case with multiple disparate clinical features, the authors were able to identify homozygous mutations in EFEMP1 as a likely cause for macular degeneration (though likely not for other features).
Conclusions: This study provides evidence that next-generation sequencing can have high success rates in a clinical setting, but also highlights key challenges. It further suggests that the presentation of known Mendelian conditions may be considerably broader than currently recognised.
Conflict of interest statement
Figures
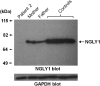
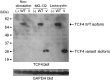
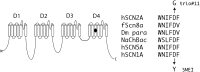
Similar articles
-
What happens when N = 1 and you want plus 1?Prenat Diagn. 2017 Jan;37(1):70-72. doi: 10.1002/pd.4975. Epub 2016 Dec 30. Prenat Diagn. 2017. PMID: 27885678
-
Clinical exome sequencing for genetic identification of rare Mendelian disorders.JAMA. 2014 Nov 12;312(18):1880-7. doi: 10.1001/jama.2014.14604. JAMA. 2014. PMID: 25326637 Free PMC article.
-
Molecular findings among patients referred for clinical whole-exome sequencing.JAMA. 2014 Nov 12;312(18):1870-9. doi: 10.1001/jama.2014.14601. JAMA. 2014. PMID: 25326635 Free PMC article.
-
Exome sequencing identifies a de novo SCN2A mutation in a patient with intractable seizures, severe intellectual disability, optic atrophy, muscular hypotonia, and brain abnormalities.Epilepsia. 2014 Apr;55(4):e25-9. doi: 10.1111/epi.12554. Epub 2014 Mar 1. Epilepsia. 2014. PMID: 24579881 Review.
-
The Rise and Rise of Exome Sequencing.Public Health Genomics. 2016;19(6):315-324. doi: 10.1159/000450991. Epub 2016 Nov 30. Public Health Genomics. 2016. PMID: 27898412 Review.
Cited by
-
N-glycoproteomics reveals distinct glycosylation alterations in NGLY1-deficient patient-derived dermal fibroblasts.J Inherit Metab Dis. 2023 Jan;46(1):76-91. doi: 10.1002/jimd.12557. Epub 2022 Oct 4. J Inherit Metab Dis. 2023. PMID: 36102038 Free PMC article.
-
Next-generation sequencing in understanding complex neurological disease.Expert Rev Neurother. 2013 Feb;13(2):215-27. doi: 10.1586/ern.12.165. Expert Rev Neurother. 2013. PMID: 23368808 Free PMC article. Review.
-
An assessment of time involved in pre-test case review and counseling for a whole genome sequencing clinical research program.J Genet Couns. 2014 Aug;23(4):516-21. doi: 10.1007/s10897-014-9697-4. Epub 2014 Feb 27. J Genet Couns. 2014. PMID: 24573557 Free PMC article.
-
Structured reviews for data and knowledge-driven research.Database (Oxford). 2020 Jan 1;2020:baaa015. doi: 10.1093/database/baaa015. Database (Oxford). 2020. PMID: 32283553 Free PMC article.
-
Protein interaction network of alternatively spliced isoforms from brain links genetic risk factors for autism.Nat Commun. 2014 Apr 11;5:3650. doi: 10.1038/ncomms4650. Nat Commun. 2014. PMID: 24722188 Free PMC article.
References
-
- Worthey EA, Mayer AN, Syverson GD, Helbling D, Bonacci BB, Decker B, Serpe JM, Dasu T, Tschannen MR, Veith RL, Basehore MJ, Broeckel U, Tomita-Mitchell A, Arca MJ, Casper JT, Margolis DA, Bick DP, Hessner MJ, Routes JM, Verbsky JW, Jacob HJ, Dimmock DP. Making a definitive diagnosis: successful clinical application of whole exome sequencing in a child with intractable inflammatory bowel disease. Genet Med 2011;13:255–62 - PubMed
-
- Vissers LE, de Ligt J, Gilissen C, Janssen I, Steehouwer M, de Vries P, van Lier B, Arts P, Wieskamp N, del Rosario M, van Bon BW, Hoischen A, de Vries BB, Brunner HG, Veltman JA. A de novo paradigm for mental retardation. Nat Genet 2010;42:1109–12 - PubMed
-
- Girard SL, Gauthier J, Noreau A, Xiong L, Zhou S, Jouan L, Dionne-Laporte A, Spiegelman D, Henrion E, Diallo O, Thibodeau P, Bachand I, Bao JY, Tong AH, Lin CH, Millet B, Jaafari N, Joober R, Dion PA, Lok S, Krebs MO, Rouleau GA. Increased exonic de novo mutation rate in individuals with schizophrenia. Nat Genet 2011;43:860–3 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous
