mGluR5 and NMDA receptors drive the experience- and activity-dependent NMDA receptor NR2B to NR2A subunit switch
- PMID: 21521618
- PMCID: PMC3087383
- DOI: 10.1016/j.neuron.2011.02.045
mGluR5 and NMDA receptors drive the experience- and activity-dependent NMDA receptor NR2B to NR2A subunit switch
Abstract
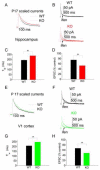
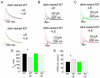
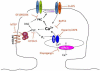
In cerebral cortex there is a developmental switch from NR2B- to NR2A-containing NMDA receptors (NMDARs) driven by activity and sensory experience. This subunit switch alters NMDAR function, influences synaptic plasticity, and its dysregulation is associated with neurological disorders. However, the mechanisms driving the subunit switch are not known. Here, we show in hippocampal CA1 pyramidal neurons that the NR2B to NR2A switch driven acutely by activity requires activation of NMDARs and mGluR5, involves PLC, Ca(2+) release from IP(3)R-dependent stores, and PKC activity. In mGluR5 knockout mice the developmental NR2B-NR2A switch in CA1 is deficient. Moreover, in visual cortex of mGluR5 knockout mice, the NR2B-NR2A switch evoked in vivo by visual experience is absent. Thus, we establish that mGluR5 and NMDARs are required for the activity-dependent NR2B-NR2A switch and play a critical role in experience-dependent regulation of NMDAR subunit composition in vivo.
Copyright © 2011 Elsevier Inc. All rights reserved.
Figures

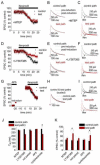
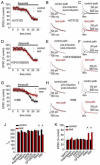
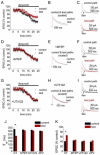
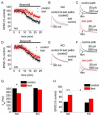
Similar articles
-
Selective subunit antagonists suggest an inhibitory relationship between NR2B and NR2A-subunit containing N-methyl-D: -aspartate receptors in hippocampal slices.Exp Brain Res. 2005 Apr;162(3):374-83. doi: 10.1007/s00221-004-2193-6. Epub 2004 Dec 3. Exp Brain Res. 2005. PMID: 15580338
-
Casein kinase 2 regulates the NR2 subunit composition of synaptic NMDA receptors.Neuron. 2010 Sep 23;67(6):984-96. doi: 10.1016/j.neuron.2010.08.011. Neuron. 2010. PMID: 20869595 Free PMC article.
-
Therapeutic significance of NR2B-containing NMDA receptors and mGluR5 metabotropic glutamate receptors in mediating the synaptotoxic effects of β-amyloid oligomers on long-term potentiation (LTP) in murine hippocampal slices.Neuropharmacology. 2011 May;60(6):982-90. doi: 10.1016/j.neuropharm.2011.01.051. Epub 2011 Feb 12. Neuropharmacology. 2011. PMID: 21310164
-
Differential roles of NR2A and NR2B-containing NMDA receptors in LTP and LTD in the CA1 region of two-week old rat hippocampus.Neuropharmacology. 2007 Jan;52(1):60-70. doi: 10.1016/j.neuropharm.2006.07.013. Epub 2006 Aug 10. Neuropharmacology. 2007. PMID: 16904707
-
Hippocampal long-term synaptic plasticity and signal amplification of NMDA receptors.Crit Rev Neurobiol. 2006;18(1-2):71-84. doi: 10.1615/critrevneurobiol.v18.i1-2.80. Crit Rev Neurobiol. 2006. PMID: 17725510 Review.
Cited by
-
Neural Influences on Tumor Progression Within the Central Nervous System.CNS Neurosci Ther. 2024 Oct;30(10):e70097. doi: 10.1111/cns.70097. CNS Neurosci Ther. 2024. PMID: 39469896 Free PMC article. Review.
-
Short-term field stimulation mimics synaptic maturation of hippocampal synapses.J Physiol. 2012 Apr 1;590(7):1641-54. doi: 10.1113/jphysiol.2011.224964. Epub 2012 Feb 20. J Physiol. 2012. PMID: 22351628 Free PMC article.
-
Early-life lead exposure recapitulates the selective loss of parvalbumin-positive GABAergic interneurons and subcortical dopamine system hyperactivity present in schizophrenia.Transl Psychiatry. 2015 Mar 10;5(3):e522. doi: 10.1038/tp.2014.147. Transl Psychiatry. 2015. PMID: 25756805 Free PMC article.
-
Effects of sex and chronic neonatal nicotine treatment on Na²⁺/K⁺/Cl⁻ co-transporter 1, K⁺/Cl⁻ co-transporter 2, brain-derived neurotrophic factor, NMDA receptor subunit 2A and NMDA receptor subunit 2B mRNA expression in the postnatal rat hippocampus.Neuroscience. 2012 Dec 6;225:105-17. doi: 10.1016/j.neuroscience.2012.09.002. Epub 2012 Sep 12. Neuroscience. 2012. PMID: 22982626 Free PMC article.
-
Synaptic plasticity of NMDA receptors: mechanisms and functional implications.Curr Opin Neurobiol. 2012 Jun;22(3):496-508. doi: 10.1016/j.conb.2012.01.007. Epub 2012 Feb 8. Curr Opin Neurobiol. 2012. PMID: 22325859 Free PMC article. Review.
References
-
- Barria A, Malinow R. Subunit-specific NMDA receptor trafficking to synapses. Neuron. 2002;35:345–353. - PubMed
-
- Barria A, Malinow R. NMDA receptor subunit composition controls synaptic plasticity by regulating binding to CaMKII. Neuron. 2005;48:289–301. - PubMed
-
- Bartlett TE, Bannister NJ, Collett VJ, Dargan SL, Massey PV, Bortolotto ZA, Fitzjohn SM, Bashir ZI, Collingridge GL, Lodge D. Differential roles of NR2A and NR2B-containing NMDA receptors in LTP and LTD in the CA1 region of two-week old rat hippocampus. Neuropharmacology. 2007;52:60–70. - PubMed
-
- Bashir ZI, Alford S, Davies SN, Randall AD, Collingridge GL. Long-term potentiation of NMDA receptor-mediated synaptic transmission in the hippocampus. Nature. 1991;349:156–158. - PubMed
-
- Bellone C, Nicoll RA. Rapid bidirectional switching of synaptic NMDA receptors. Neuron. 2007;55:779–785. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

