Recurrent rearrangements of chromosome 1q21.1 and variable pediatric phenotypes
- PMID: 18784092
- PMCID: PMC2703742
- DOI: 10.1056/NEJMoa0805384
Recurrent rearrangements of chromosome 1q21.1 and variable pediatric phenotypes
Abstract
Background: Duplications and deletions in the human genome can cause disease or predispose persons to disease. Advances in technologies to detect these changes allow for the routine identification of submicroscopic imbalances in large numbers of patients.
Methods: We tested for the presence of microdeletions and microduplications at a specific region of chromosome 1q21.1 in two groups of patients with unexplained mental retardation, autism, or congenital anomalies and in unaffected persons.
Results: We identified 25 persons with a recurrent 1.35-Mb deletion within 1q21.1 from screening 5218 patients. The microdeletions had arisen de novo in eight patients, were inherited from a mildly affected parent in three patients, were inherited from an apparently unaffected parent in six patients, and were of unknown inheritance in eight patients. The deletion was absent in a series of 4737 control persons (P=1.1x10(-7)). We found considerable variability in the level of phenotypic expression of the microdeletion; phenotypes included mild-to-moderate mental retardation, microcephaly, cardiac abnormalities, and cataracts. The reciprocal duplication was enriched in nine children with mental retardation or autism spectrum disorder and other variable features (P=0.02). We identified three deletions and three duplications of the 1q21.1 region in an independent sample of 788 patients with mental retardation and congenital anomalies.
Conclusions: We have identified recurrent molecular lesions that elude syndromic classification and whose disease manifestations must be considered in a broader context of development as opposed to being assigned to a specific disease. Clinical diagnosis in patients with these lesions may be most readily achieved on the basis of genotype rather than phenotype.
2008 Massachusetts Medical Society
Figures
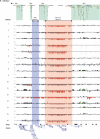

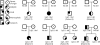

Comment in
-
Cytogenetic technology--genotype and phenotype.N Engl J Med. 2008 Oct 16;359(16):1728-30. doi: 10.1056/NEJMe0806570. Epub 2008 Sep 10. N Engl J Med. 2008. PMID: 18784093 No abstract available.
-
How best to use CGH arrays in the clinical setting.Genet Med. 2009 May;11(5):371; author reply 371-2. doi: 10.1097/gim.0b013e31819dbfb4. Genet Med. 2009. PMID: 19346958 No abstract available.
Similar articles
-
Recurrent reciprocal 16p11.2 rearrangements associated with global developmental delay, behavioural problems, dysmorphism, epilepsy, and abnormal head size.J Med Genet. 2010 May;47(5):332-41. doi: 10.1136/jmg.2009.073015. Epub 2009 Nov 12. J Med Genet. 2010. PMID: 19914906 Free PMC article.
-
Recurrent reciprocal deletions and duplications of 16p13.11: the deletion is a risk factor for MR/MCA while the duplication may be a rare benign variant.J Med Genet. 2009 Apr;46(4):223-32. doi: 10.1136/jmg.2007.055202. Epub 2008 Jun 11. J Med Genet. 2009. PMID: 18550696 Free PMC article.
-
An eight-case 1q21 region series: novel aberrations and clinical variability with new features.J Intellect Disabil Res. 2019 Jun;63(6):548-557. doi: 10.1111/jir.12592. Epub 2019 Feb 18. J Intellect Disabil Res. 2019. PMID: 30773728
-
Recurrent 1q21.1 deletion syndrome: report on variable expression, nonpenetrance and review of literature.Clin Dysmorphol. 2020 Jul;29(3):127-131. doi: 10.1097/MCD.0000000000000327. Clin Dysmorphol. 2020. PMID: 32459673 Free PMC article. Review.
-
Chromosomal phenotypes and submicroscopic abnormalities.Hum Genomics. 2004 Jan;1(2):126-33. doi: 10.1186/1479-7364-1-2-126. Hum Genomics. 2004. PMID: 15601540 Free PMC article. Review.
Cited by
-
Discovery of Rare Mutations in Autism: Elucidating Neurodevelopmental Mechanisms.Neurotherapeutics. 2015 Jul;12(3):553-71. doi: 10.1007/s13311-015-0363-9. Neurotherapeutics. 2015. PMID: 26105128 Free PMC article. Review.
-
Association testing of copy number variants in schizophrenia and autism spectrum disorders.J Neurodev Disord. 2012 May 30;4(1):15. doi: 10.1186/1866-1955-4-15. J Neurodev Disord. 2012. PMID: 22958593 Free PMC article.
-
"What does it mean?": uncertainties in understanding results of chromosomal microarray testing.Genet Med. 2012 Feb;14(2):250-8. doi: 10.1038/gim.2011.52. Epub 2012 Jan 5. Genet Med. 2012. PMID: 22241091 Free PMC article.
-
Rare copy number variants contribute to congenital left-sided heart disease.PLoS Genet. 2012 Sep;8(9):e1002903. doi: 10.1371/journal.pgen.1002903. Epub 2012 Sep 6. PLoS Genet. 2012. PMID: 22969434 Free PMC article.
-
Replicated linear association between DUF1220 copy number and severity of social impairment in autism.Hum Genet. 2015 Jun;134(6):569-75. doi: 10.1007/s00439-015-1537-6. Epub 2015 Mar 11. Hum Genet. 2015. PMID: 25758905 Free PMC article.
References
-
- Barber JC, Maloney VK, Huang S, et al. 8p23.1 Duplication syndrome: a novel genomic condition with unexpected complexity revealed by array CGH. Eur J Hum Genet. 2008;16:18–27. - PubMed
-
- Koolen DA, Vissers LE, Pfundt R, et al. A new chromosome 17q21.31 microdeletion syndrome associated with a common inversion polymorphism. Nat Genet. 2006;38:999–1001. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials
Miscellaneous
