Recurrent reciprocal genomic rearrangements of 17q12 are associated with renal disease, diabetes, and epilepsy
- PMID: 17924346
- PMCID: PMC2265663
- DOI: 10.1086/522591
Recurrent reciprocal genomic rearrangements of 17q12 are associated with renal disease, diabetes, and epilepsy
Abstract
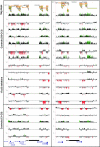
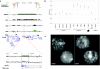
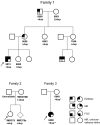
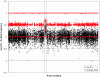
Most studies of genomic disorders have focused on patients with cognitive disability and/or peripheral nervous system defects. In an effort to broaden the phenotypic spectrum of this disease model, we assessed 155 autopsy samples from fetuses with well-defined developmental pathologies in regions predisposed to recurrent rearrangement, by array-based comparative genomic hybridization. We found that 6% of fetal material showed evidence of microdeletion or microduplication, including three independent events that likely resulted from unequal crossing-over between segmental duplications. One of the microdeletions, identified in a fetus with multicystic dysplastic kidneys, encompasses the TCF2 gene on 17q12, previously shown to be mutated in maturity-onset diabetes, as well as in a subset of pediatric renal abnormalities. Fine-scale mapping of the breakpoints in different patient cohorts revealed a recurrent 1.5-Mb de novo deletion in individuals with phenotypes that ranged from congenital renal abnormalities to maturity-onset diabetes of the young type 5. We also identified the reciprocal duplication, which appears to be enriched in samples from patients with epilepsy. We describe the first example of a recurrent genomic disorder associated with diabetes.
Figures



Similar articles
-
Clinical spectrum associated with recurrent genomic rearrangements in chromosome 17q12.Eur J Hum Genet. 2010 Mar;18(3):278-84. doi: 10.1038/ejhg.2009.174. Epub 2009 Oct 21. Eur J Hum Genet. 2010. PMID: 19844256 Free PMC article.
-
Discovery of previously unidentified genomic disorders from the duplication architecture of the human genome.Nat Genet. 2006 Sep;38(9):1038-42. doi: 10.1038/ng1862. Epub 2006 Aug 13. Nat Genet. 2006. PMID: 16906162
-
Chromosome 17q12 duplications: Further delineation of the range of psychiatric and clinical phenotypes.Am J Med Genet B Neuropsychiatr Genet. 2018 Jul;177(5):520-528. doi: 10.1002/ajmg.b.32643. Am J Med Genet B Neuropsychiatr Genet. 2018. PMID: 30134084
-
Duplication hotspots, rare genomic disorders, and common disease.Curr Opin Genet Dev. 2009 Jun;19(3):196-204. doi: 10.1016/j.gde.2009.04.003. Epub 2009 May 22. Curr Opin Genet Dev. 2009. PMID: 19477115 Free PMC article. Review.
-
17q12 deletion and duplication syndrome in Denmark-A clinical cohort of 38 patients and review of the literature.Am J Med Genet A. 2016 Nov;170(11):2934-2942. doi: 10.1002/ajmg.a.37848. Epub 2016 Jul 13. Am J Med Genet A. 2016. PMID: 27409573 Review.
Cited by
-
Exome Sequencing in Fetuses with Structural Malformations.J Clin Med. 2014 Jul 8;3(3):747-62. doi: 10.3390/jcm3030747. J Clin Med. 2014. PMID: 26237476 Free PMC article. Review.
-
Prenatal diagnosis of 17q12 copy number variants in fetuses via chromosomal microarray analysis - A retrospective cohort study and literature review.Heliyon. 2024 Aug 19;10(17):e36558. doi: 10.1016/j.heliyon.2024.e36558. eCollection 2024 Sep 15. Heliyon. 2024. PMID: 39286125 Free PMC article.
-
Genetic testing in renal disease.Pediatr Nephrol. 2012 Jun;27(6):873-83. doi: 10.1007/s00467-011-1865-2. Epub 2011 May 27. Pediatr Nephrol. 2012. PMID: 21617915 Review.
-
Identification of a heterozygous genomic deletion in the spatacsin gene in SPG11 patients using high-resolution comparative genomic hybridization.Neurogenetics. 2009 Feb;10(1):43-8. doi: 10.1007/s10048-008-0144-2. Epub 2008 Sep 12. Neurogenetics. 2009. PMID: 18787847
-
Human copy number variation and complex genetic disease.Annu Rev Genet. 2011;45:203-26. doi: 10.1146/annurev-genet-102209-163544. Epub 2011 Aug 19. Annu Rev Genet. 2011. PMID: 21854229 Free PMC article.
References
Web Resources
-
- Human Genome Structural Variation Project, http://humanparalogy.gs.washington.edu/structuralvariation/ (for oligonucleotide array data)
-
- Online Mendelian Inheritance in Man (OMIM), http://www.ncbi.nlm.nih.gov/Omim/ (for MODY5, CCL3L1, CCL4L1, TBC1D3, TCF2, and LHX1)
-
- PharmGKB: PARC Profile, http://www.pharmgkb.org/network/members/parc.jsp#team
References
-
- Menten B, Maas N, Thienpont B, Buysse K, Vandesompele J, Melotte C, de Ravel T, Van Vooren S, Balikova I, Backx L, et al (2006) Emerging patterns of cryptic chromosomal imbalance in patients with idiopathic mental retardation and multiple congenital anomalies: a new series of 140 patients and review of published reports. J Med Genet 43:625–63310.1136/jmg.2005.039453 - DOI - PMC - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Miscellaneous

