NFIA haploinsufficiency is associated with a CNS malformation syndrome and urinary tract defects
- PMID: 17530927
- PMCID: PMC1877820
- DOI: 10.1371/journal.pgen.0030080
NFIA haploinsufficiency is associated with a CNS malformation syndrome and urinary tract defects
Abstract
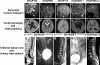
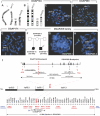
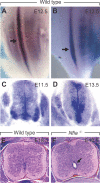
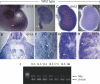
Complex central nervous system (CNS) malformations frequently coexist with other developmental abnormalities, but whether the associated defects share a common genetic basis is often unclear. We describe five individuals who share phenotypically related CNS malformations and in some cases urinary tract defects, and also haploinsufficiency for the NFIA transcription factor gene due to chromosomal translocation or deletion. Two individuals have balanced translocations that disrupt NFIA. A third individual and two half-siblings in an unrelated family have interstitial microdeletions that include NFIA. All five individuals exhibit similar CNS malformations consisting of a thin, hypoplastic, or absent corpus callosum, and hydrocephalus or ventriculomegaly. The majority of these individuals also exhibit Chiari type I malformation, tethered spinal cord, and urinary tract defects that include vesicoureteral reflux. Other genes are also broken or deleted in all five individuals, and may contribute to the phenotype. However, the only common genetic defect is NFIA haploinsufficiency. In addition, previous analyses of Nfia(-/-) knockout mice indicate that Nfia deficiency also results in hydrocephalus and agenesis of the corpus callosum. Further investigation of the mouse Nfia(+/-) and Nfia(-/-) phenotypes now reveals that, at reduced penetrance, Nfia is also required in a dosage-sensitive manner for ureteral and renal development. Nfia is expressed in the developing ureter and metanephric mesenchyme, and Nfia(+/-) and Nfia(-/-) mice exhibit abnormalities of the ureteropelvic and ureterovesical junctions, as well as bifid and megaureter. Collectively, the mouse Nfia mutant phenotype and the common features among these five human cases indicate that NFIA haploinsufficiency contributes to a novel human CNS malformation syndrome that can also include ureteral and renal defects.
Conflict of interest statement
Competing interests. The authors have declared that no competing interests exist.
Figures


Similar articles
-
Phenotypic Spectrum of NFIA Haploinsufficiency: Two Additional Cases and Review of the Literature.Genes (Basel). 2022 Nov 30;13(12):2249. doi: 10.3390/genes13122249. Genes (Basel). 2022. PMID: 36553517 Free PMC article. Review.
-
Loss-of-function variants in NFIA provide further support that NFIA is a critical gene in 1p32-p31 deletion syndrome: A four patient series.Am J Med Genet A. 2017 Dec;173(12):3158-3164. doi: 10.1002/ajmg.a.38460. Epub 2017 Sep 22. Am J Med Genet A. 2017. PMID: 28941020
-
An intragenic deletion of the NFIA gene in a patient with a hypoplastic corpus callosum, craniofacial abnormalities and urinary tract defects.Eur J Med Genet. 2014 Feb;57(2-3):65-70. doi: 10.1016/j.ejmg.2013.12.011. Epub 2014 Jan 22. Eur J Med Genet. 2014. PMID: 24462883
-
Chromosome 1p32-p31 deletion syndrome: prenatal diagnosis by array comparative genomic hybridization using uncultured amniocytes and association with NFIA haploinsufficiency, ventriculomegaly, corpus callosum hypogenesis, abnormal external genitalia, and intrauterine growth restriction.Taiwan J Obstet Gynecol. 2011 Sep;50(3):345-52. doi: 10.1016/j.tjog.2011.07.014. Taiwan J Obstet Gynecol. 2011. PMID: 22030051
-
Variants in nuclear factor I genes influence growth and development.Am J Med Genet C Semin Med Genet. 2019 Dec;181(4):611-626. doi: 10.1002/ajmg.c.31747. Epub 2019 Nov 15. Am J Med Genet C Semin Med Genet. 2019. PMID: 31730271 Review.
Cited by
-
Lower urinary tract development and disease.Wiley Interdiscip Rev Syst Biol Med. 2013 May-Jun;5(3):307-42. doi: 10.1002/wsbm.1212. Epub 2013 Feb 13. Wiley Interdiscip Rev Syst Biol Med. 2013. PMID: 23408557 Free PMC article. Review.
-
Truncating mutation in NFIA causes brain malformation and urinary tract defects.Hum Genome Var. 2015 Feb 26;2:15007. doi: 10.1038/hgv.2015.7. eCollection 2015. Hum Genome Var. 2015. PMID: 27081522 Free PMC article.
-
Phenotypic Spectrum of NFIA Haploinsufficiency: Two Additional Cases and Review of the Literature.Genes (Basel). 2022 Nov 30;13(12):2249. doi: 10.3390/genes13122249. Genes (Basel). 2022. PMID: 36553517 Free PMC article. Review.
-
Cell-specific NFIA upregulation promotes epileptogenesis by TRPV4-mediated astrocyte reactivity.J Neuroinflammation. 2023 Oct 25;20(1):247. doi: 10.1186/s12974-023-02909-4. J Neuroinflammation. 2023. PMID: 37880726 Free PMC article.
-
De novo variants in SUPT16H cause neurodevelopmental disorders associated with corpus callosum abnormalities.J Med Genet. 2020 Jul;57(7):461-465. doi: 10.1136/jmedgenet-2019-106193. Epub 2020 Jan 10. J Med Genet. 2020. PMID: 31924697 Free PMC article.
References
-
- Kleinjan DJ, van Heyningen V. Position effect in human genetic disease. Hum Mol Genet. 1998;7:1611–1618. - PubMed
-
- Guillem P, Fabre B, Cans C, Robert-Gnansia E, Jouk PS. Trends in elective terminations of pregnancy between 1989 and 2000 in a French county (the Isere) Prenat Diagn. 2003;23:877–883. - PubMed
-
- Paul LK, Brown WS, Adolphs R, Tyszka JM, Richards LJ, et al. Agenesis of the corpus callosum: genetic, developmental and functional aspects of connectivity. Nat Rev Neurosci. 2007;8:287–299. - PubMed
-
- Wang LW, Huang CC, Yeh TF. Major brain lesions detected on sonographic screening of apparently normal term neonates. Neuroradiology. 2004;46:368–373. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

