Structural and kinetic basis for substrate selectivity in Populus tremuloides sinapyl alcohol dehydrogenase
- PMID: 15829607
- PMCID: PMC1091777
- DOI: 10.1105/tpc.104.029983
Structural and kinetic basis for substrate selectivity in Populus tremuloides sinapyl alcohol dehydrogenase
Abstract
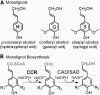
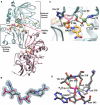
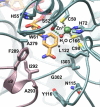
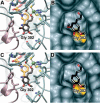
We describe the three-dimensional structure of sinapyl alcohol dehydrogenase (SAD) from Populus tremuloides (aspen), a member of the NADP(H)-dependent dehydrogenase family that catalyzes the last reductive step in the formation of monolignols. The active site topology revealed by the crystal structure substantiates kinetic results indicating that SAD maintains highest specificity for the substrate sinapaldehyde. We also report substantial substrate inhibition kinetics for the SAD-catalyzed reduction of hydroxycinnamaldehydes. Although SAD and classical cinnamyl alcohol dehydrogenases (CADs) catalyze the same reaction and share some sequence identity, the active site topology of SAD is strikingly different from that predicted for classical CADs. Kinetic analyses of wild-type SAD and several active site mutants demonstrate the complexity of defining determinants of substrate specificity in these enzymes. These results, along with a phylogenetic analysis, support the inclusion of SAD in a plant alcohol dehydrogenase subfamily that includes cinnamaldehyde and benzaldehyde dehydrogenases. We used the SAD three-dimensional structure to model several of these SAD-like enzymes, and although their active site topologies largely mirror that of SAD, we describe a correlation between substrate specificity and amino acid substitution patterns in their active sites. The SAD structure thus provides a framework for understanding substrate specificity in this family of enzymes and for engineering new enzyme specificities.
Figures
Similar articles
-
The last step of syringyl monolignol biosynthesis in angiosperms is regulated by a novel gene encoding sinapyl alcohol dehydrogenase.Plant Cell. 2001 Jul;13(7):1567-86. doi: 10.1105/tpc.010111. Plant Cell. 2001. PMID: 11449052 Free PMC article.
-
Syringyl lignin is unaltered by severe sinapyl alcohol dehydrogenase suppression in tobacco.Plant Cell. 2011 Dec;23(12):4492-506. doi: 10.1105/tpc.111.089037. Epub 2011 Dec 9. Plant Cell. 2011. PMID: 22158465 Free PMC article.
-
The Enzyme Activity and Substrate Specificity of Two Major Cinnamyl Alcohol Dehydrogenases in Sorghum (Sorghum bicolor), SbCAD2 and SbCAD4.Plant Physiol. 2017 Aug;174(4):2128-2145. doi: 10.1104/pp.17.00576. Epub 2017 Jun 12. Plant Physiol. 2017. PMID: 28606901 Free PMC article.
-
Kinetic interpretations of active site topologies and residue exchanges in Drosophila alcohol dehydrogenases.Int J Biochem. 1992 Feb;24(2):169-81. doi: 10.1016/0020-711x(92)90245-v. Int J Biochem. 1992. PMID: 1733784 Review.
-
Trends in lignin modification: a comprehensive analysis of the effects of genetic manipulations/mutations on lignification and vascular integrity.Phytochemistry. 2002 Oct;61(3):221-94. doi: 10.1016/s0031-9422(02)00211-x. Phytochemistry. 2002. PMID: 12359514 Review.
Cited by
-
Transcriptome Mining Provides Insights into Cell Wall Metabolism and Fiber Lignification in Agave tequilana Weber.Plants (Basel). 2022 Jun 2;11(11):1496. doi: 10.3390/plants11111496. Plants (Basel). 2022. PMID: 35684270 Free PMC article.
-
Recombinant Penicillium oxalicum 16 β-Glucosidase 1 Displays Comprehensive Inhibitory Resistance to Several Lignocellulose Pretreatment Products, Ethanol, and Salt.Appl Biochem Biotechnol. 2020 Jun;191(2):772-784. doi: 10.1007/s12010-019-03183-y. Epub 2019 Dec 20. Appl Biochem Biotechnol. 2020. PMID: 31858406 Free PMC article.
-
A genomewide analysis of the cinnamyl alcohol dehydrogenase family in sorghum [Sorghum bicolor (L.) Moench] identifies SbCAD2 as the brown midrib6 gene.Genetics. 2009 Feb;181(2):783-95. doi: 10.1534/genetics.108.098996. Epub 2008 Dec 15. Genetics. 2009. PMID: 19087955 Free PMC article.
-
Nucleotide sequence of Phaseolus vulgaris L. alcohol dehydrogenase encoding cDNA and three-dimensional structure prediction of the deduced protein.Pharmacognosy Res. 2015 Apr-Jun;7(2):203-8. doi: 10.4103/0974-8490.150532. Pharmacognosy Res. 2015. PMID: 25829796 Free PMC article.
-
Manipulation of Guaiacyl and Syringyl Monomer Biosynthesis in an Arabidopsis Cinnamyl Alcohol Dehydrogenase Mutant Results in Atypical Lignin Biosynthesis and Modified Cell Wall Structure.Plant Cell. 2015 Aug;27(8):2195-209. doi: 10.1105/tpc.15.00373. Epub 2015 Aug 11. Plant Cell. 2015. PMID: 26265762 Free PMC article.
References
-
- Baker, P.J., Britton, K.L., Rice, D.W., Rob, A., and Stillman, T.J. (1992). Structural consequences of sequence patterns in the fingerprint region of the nucleotide binding fold: Implications for nucleotide specificity. J. Mol. Biol. 228, 662–671. - PubMed
-
- Banfield, M.J., Salvucci, M.E., Baker, E.N., and Smith, C.A. (2001). Crystal structure of the NADP(H)-dependent ketose reductase from Bemisia argentifolii at 2.3 A resolution. J. Mol. Biol. 306, 239–250. - PubMed
-
- Blanco-Portales, R., Medina-Escobar, N., Lopez-Raez, J.A., Gonzalez-Reyes, J.A., Villalba, J.M., Moyano, E., Caballero, J.L., and Munoz-Blanco, J. (2002). Cloning, expression and immunolocalization pattern of a cinnamyl alcohol dehydrogenase gene from strawberry (Fragaria × ananassa cv. Chandler). J. Exp. Bot. 53, 1723–1734. - PubMed
-
- Brill, E.M., Abrahams, S., Hayes, C.M., Jenkins, C.L., and Watson, J.M. (1999). Molecular characterisation and expression of a wound-inducible cDNA encoding a novel cinnamyl-alcohol dehydrogenase enzyme in lucerne (Medicago sativa L.). Plant Mol. Biol. 41, 279–291. - PubMed
-
- Brunger, A.T., et al. (1998). Crystallography & NMR system: A new software suite for macromolecular structure determination. Acta Crystallogr. D Biol. Crystallogr. 54, 905–921. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

