Complete absence of Cockayne syndrome group B gene product gives rise to UV-sensitive syndrome but not Cockayne syndrome
- PMID: 15486090
- PMCID: PMC524447
- DOI: 10.1073/pnas.0404587101
Complete absence of Cockayne syndrome group B gene product gives rise to UV-sensitive syndrome but not Cockayne syndrome
Abstract
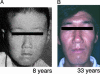
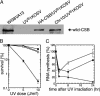
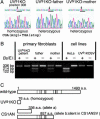
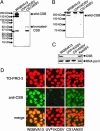
UV-sensitive syndrome (UVsS) is a rare autosomal recessive disorder characterized by photosensitivity and mild freckling but without neurological abnormalities or skin tumors. UVsS cells show UV hypersensitivity and defective transcription-coupled DNA repair of UV damage. It was suggested that UVsS does not belong to any complementation groups of known photosensitive disorders such as xeroderma pigmentosum and Cockayne syndrome (CS). To identify the gene responsible for UVsS, we performed a microcell-mediated chromosome transfer based on the functional complementation of UV hypersensitivity. We found that one of the UVsS cell lines, UVs1KO, acquired UV resistance when human chromosome 10 was transferred. Because the gene responsible for CS group B (CSB), which involves neurological abnormalities and photosensitivity as well as a defect in transcription-coupled DNA repair of UV damage, is located on chromosome 10, we sequenced the CSB gene from UVs1KO and detected a homozygous null mutation. Our results indicate that previous complementation analysis of UVs1KO was erroneous. This finding was surprising because a null mutation of the CSB gene would be expected to result in CS features such as severe developmental and neurological abnormalities. On the other hand, no mutation in the CSB cDNA and a normal amount of CSB protein was detected in Kps3, a UVsS cell line obtained from an unrelated patient, indicating genetic heterogeneity in UVsS. Possible explanations for the discrepancy in the genotype-phenotype relationship in UVs1KO are presented.
Figures
Comment in
-
The many faces of Cockayne syndrome.Proc Natl Acad Sci U S A. 2004 Oct 26;101(43):15273-4. doi: 10.1073/pnas.0406894101. Epub 2004 Oct 19. Proc Natl Acad Sci U S A. 2004. PMID: 15494443 Free PMC article. No abstract available.
Similar articles
-
Differential requirement for the ATPase domain of the Cockayne syndrome group B gene in the processing of UV-induced DNA damage and 8-oxoguanine lesions in human cells.Nucleic Acids Res. 2002 Feb 1;30(3):782-93. doi: 10.1093/nar/30.3.782. Nucleic Acids Res. 2002. PMID: 11809892 Free PMC article.
-
Alterations in the CSB gene in three Italian patients with the severe form of Cockayne syndrome (CS) but without clinical photosensitivity.Hum Mol Genet. 1999 May;8(5):935-41. doi: 10.1093/hmg/8.5.935. Hum Mol Genet. 1999. PMID: 10196384
-
Role of the ATPase domain of the Cockayne syndrome group B protein in UV induced apoptosis.Oncogene. 2000 Jan 27;19(4):477-89. doi: 10.1038/sj.onc.1203372. Oncogene. 2000. PMID: 10698517
-
The role of Cockayne Syndrome group B (CSB) protein in base excision repair and aging.Mech Ageing Dev. 2008 Jul-Aug;129(7-8):441-8. doi: 10.1016/j.mad.2008.04.009. Epub 2008 Apr 30. Mech Ageing Dev. 2008. PMID: 18541289 Free PMC article. Review.
-
Cockayne syndrome group B cellular and biochemical functions.Am J Hum Genet. 2003 Dec;73(6):1217-39. doi: 10.1086/380399. Epub 2003 Nov 24. Am J Hum Genet. 2003. PMID: 14639525 Free PMC article. Review.
Cited by
-
UVSSA and USP7, a new couple in transcription-coupled DNA repair.Chromosoma. 2013 Aug;122(4):275-84. doi: 10.1007/s00412-013-0420-2. Epub 2013 Jun 13. Chromosoma. 2013. PMID: 23760561 Free PMC article. Review.
-
DDB2 complex-mediated ubiquitylation around DNA damage is oppositely regulated by XPC and Ku and contributes to the recruitment of XPA.Mol Cell Biol. 2010 Jun;30(11):2708-23. doi: 10.1128/MCB.01460-09. Epub 2010 Apr 5. Mol Cell Biol. 2010. PMID: 20368362 Free PMC article.
-
DNA repair: dynamic defenders against cancer and aging.PLoS Biol. 2006 Jun;4(6):e203. doi: 10.1371/journal.pbio.0040203. Epub 2006 Jun 13. PLoS Biol. 2006. PMID: 16752948 Free PMC article.
-
Cockayne syndrome in adults: review with clinical and pathologic study of a new case.J Child Neurol. 2006 Nov;21(11):991-1006. doi: 10.1177/08830738060210110101. J Child Neurol. 2006. PMID: 17092472 Free PMC article. Review.
-
RNA polymerase between lesion bypass and DNA repair.Cell Mol Life Sci. 2013 Dec;70(23):4495-509. doi: 10.1007/s00018-013-1384-3. Epub 2013 Jun 27. Cell Mol Life Sci. 2013. PMID: 23807206 Free PMC article. Review.
References
-
- Friedberg, E. C., Walker, G. C. & Sied, W. (1995) DNA Repair and Mutagenesis (Am. Soc. Microbiol., Washington, DC).
-
- Hanawalt, P. C. (2002) Oncogene 21, 8949-8956. - PubMed
-
- Masutani, C., Kusumoto, R., Yamada, A., Dohmae, N., Yokoi, M., Yuasa, M., Araki, M., Iwai, S., Takio, K. & Hanaoka, F. (1999) Nature 399, 700-704. - PubMed
-
- Nance, M. A. & Berry, S. A. (1992) Am. J. Med. Genet. 42, 68-84. - PubMed
-
- Mayne, L. V. & Lehmann, A. R. (1982) Cancer Res. 42, 1473-1478. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials

