N-terminus of the rat adenine glycosylase MYH affects excision rates and processing of MYH-generated abasic sites
- PMID: 15310837
- PMCID: PMC514379
- DOI: 10.1093/nar/gkh758
N-terminus of the rat adenine glycosylase MYH affects excision rates and processing of MYH-generated abasic sites
Abstract
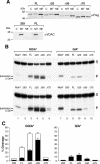
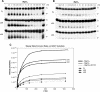
Repair of most modified and mispaired bases in the genome is initiated by DNA glycosylases, which bind to their respective targets and cleave the N-glycosyl bond to initiate base excision repair (BER). The mammalian homolog of the Escherichia coli MutY DNA glycosylase (MYH) cleaves adenine residues paired with either oxidized or non-modified guanines. MYH is crucial for the avoidance of mutations resulting from oxidative DNA damage. Multiple N-terminal splice variants of MYH exist in mammalian cells and it is likely that different variants result in the production of enzymes with altered properties. To investigate whether modifications in the N-terminus are consequential to MYH function, we overexpressed intact and N-terminal-deletion rat MYH proteins and examined their activities. We found that deletion of 75 amino acids, which perturbs the catalytic core that is conserved with E.coli MutY, abolished excision activity. In contrast, deletions limited to the extended mammalian N-terminal domain, differentially influenced steady-state excision rates. Notably, deletion of 50 amino acids resulted in an enzyme with a significantly lower K(m) favoring formation of excision products with 3'-OH termini. Our findings suggest that MYH isoforms divergent in the N-terminus influence excision rates and processing of abasic sites.
Figures
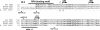


Similar articles
-
Isolation and analyses of MutY homologs (MYH).Methods Enzymol. 2006;408:64-78. doi: 10.1016/S0076-6879(06)08005-0. Methods Enzymol. 2006. PMID: 16793363
-
Replication-associated repair of adenine:8-oxoguanine mispairs by MYH.Curr Biol. 2002 Feb 19;12(4):335-9. doi: 10.1016/s0960-9822(02)00686-3. Curr Biol. 2002. PMID: 11864576
-
The C-terminal domain of the adenine-DNA glycosylase MutY confers specificity for 8-oxoguanine.adenine mispairs and may have evolved from MutT, an 8-oxo-dGTPase.Biochemistry. 1999 May 18;38(20):6374-9. doi: 10.1021/bi990335x. Biochemistry. 1999. PMID: 10350454
-
Oxidative DNA damage repair in mammalian cells: a new perspective.DNA Repair (Amst). 2007 Apr 1;6(4):470-80. doi: 10.1016/j.dnarep.2006.10.011. Epub 2006 Nov 20. DNA Repair (Amst). 2007. PMID: 17116430 Free PMC article. Review.
-
The intricate structural chemistry of base excision repair machinery: implications for DNA damage recognition, removal, and repair.DNA Repair (Amst). 2007 Apr 1;6(4):410-28. doi: 10.1016/j.dnarep.2006.10.004. Epub 2007 Jan 8. DNA Repair (Amst). 2007. PMID: 17208522 Review.
Cited by
-
Accumulation of oxidatively generated DNA damage in the brain: a mechanism of neurotoxicity.Free Radic Biol Med. 2007 Feb 1;42(3):385-93. doi: 10.1016/j.freeradbiomed.2006.11.009. Epub 2006 Nov 10. Free Radic Biol Med. 2007. PMID: 17210451 Free PMC article.
-
Isoforms of Base Excision Repair Enzymes Produced by Alternative Splicing.Int J Mol Sci. 2019 Jul 3;20(13):3279. doi: 10.3390/ijms20133279. Int J Mol Sci. 2019. PMID: 31277343 Free PMC article. Review.
-
A haplotype variation affecting the mitochondrial transportation of hMYH protein could be a risk factor for colorectal cancer in Chinese.BMC Cancer. 2008 Sep 23;8:269. doi: 10.1186/1471-2407-8-269. BMC Cancer. 2008. PMID: 18811933 Free PMC article.
-
Elevated metals compromise repair of oxidative DNA damage via the base excision repair pathway: implications of pathologic iron overload in the brain on integrity of neuronal DNA.J Neurochem. 2009 Sep;110(6):1774-83. doi: 10.1111/j.1471-4159.2009.06271.x. Epub 2009 Jul 8. J Neurochem. 2009. PMID: 19619136 Free PMC article.
-
MUTYH DNA glycosylase: the rationale for removing undamaged bases from the DNA.Front Genet. 2013 Feb 28;4:18. doi: 10.3389/fgene.2013.00018. eCollection 2013. Front Genet. 2013. PMID: 23450852 Free PMC article.
References
-
- Beckman K.B. and Ames,B.N. (1997) Oxidative decay of DNA. J. Biol. Chem., 272, 19633–19636. - PubMed
-
- Mitra S., Hazra,T.K., Roy,R., Ikeda,S., Biswas,T., Lock,J., Boldogh,I. and Izumi,T. (1997) Complexities of DNA base excision repair in mammalian cells. Mol. Cell, 7, 305–312. - PubMed
-
- Dianov G., Bischoff,C., Piotrowski,J. and Bohr,V.A. (1998) Repair pathways for processing of 8-oxoguanine in DNA by mammalian cell extracts. J. Biol. Chem., 273, 33811–33816. - PubMed
-
- Michaels M.L., Tchou,J., Grollman,A.P. and Miller,J.H. (1992) A repair system for 8-oxo-7,8-dihydrodeoxyguanine. Biochemistry, 31, 10964–10968. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

