Multiple promoters in the WNK1 gene: one controls expression of a kidney-specific kinase-defective isoform
- PMID: 14645531
- PMCID: PMC309643
- DOI: 10.1128/MCB.23.24.9208-9221.2003
Multiple promoters in the WNK1 gene: one controls expression of a kidney-specific kinase-defective isoform
Abstract
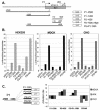
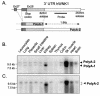
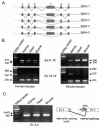
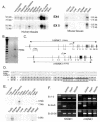
WNK1 is a serine-threonine kinase, the expression of which is affected in pseudohypoaldosteronism type II, a Mendelian form of arterial hypertension. We characterized human WNK1 transcripts to determine the molecular mechanisms governing WNK1 expression. We report the presence of two promoters generating two WNK1 isoforms with a complete kinase domain. Further variations are achieved by the use of two polyadenylation sites and tissue-specific splicing. We also determined the structure of a kidney-specific isoform regulated by a third promoter and starting at a novel exon. This transcript is kinase defective and has a predominant expression in the kidney compared to the other WNK1 isoforms, with, furthermore, a highly restricted expression profile in the distal convoluted tubule. We confirmed that the ubiquitous and kidney-specific promoters are functional in several cells lines and identified core promoters and regulatory elements. In particular, a strong enhancer element upstream from the kidney-specific exon seems specific to renal epithelial cells. Thus, control of human WNK1 gene expression of kinase-active or -deficient isoforms is mediated predominantly through the use of multiple transcription initiation sites and tissue-specific regulatory elements.
Figures



Similar articles
-
WNK1, a gene within a novel blood pressure control pathway, tissue-specifically generates radically different isoforms with and without a kinase domain.J Am Soc Nephrol. 2003 Oct;14(10):2447-56. doi: 10.1097/01.asn.0000089830.97681.3b. J Am Soc Nephrol. 2003. PMID: 14514722
-
Regulation of ROMK channel and K+ homeostasis by kidney-specific WNK1 kinase.J Biol Chem. 2009 May 1;284(18):12198-206. doi: 10.1074/jbc.M806551200. Epub 2009 Feb 25. J Biol Chem. 2009. PMID: 19244242 Free PMC article.
-
The kidney-specific WNK1 isoform is induced by aldosterone and stimulates epithelial sodium channel-mediated Na+ transport.Proc Natl Acad Sci U S A. 2004 Dec 14;101(50):17434-9. doi: 10.1073/pnas.0408146101. Epub 2004 Dec 6. Proc Natl Acad Sci U S A. 2004. PMID: 15583131 Free PMC article.
-
Pathogenesis of pseudohypoaldosteronism type 2 by WNK1 mutations.Curr Opin Nephrol Hypertens. 2012 Jan;21(1):39-45. doi: 10.1097/MNH.0b013e32834d2fde. Curr Opin Nephrol Hypertens. 2012. PMID: 22080857 Review.
-
WNK1 in the kidney.Curr Opin Nephrol Hypertens. 2022 Sep 1;31(5):471-478. doi: 10.1097/MNH.0000000000000820. Epub 2022 Jul 15. Curr Opin Nephrol Hypertens. 2022. PMID: 35894282 Review.
Cited by
-
Molecular physiology of SPAK and OSR1: two Ste20-related protein kinases regulating ion transport.Physiol Rev. 2012 Oct;92(4):1577-617. doi: 10.1152/physrev.00009.2012. Physiol Rev. 2012. PMID: 23073627 Free PMC article. Review.
-
Comparative analysis of the expression profile of Wnk1 and Wnk1/Hsn2 splice variants in developing and adult mouse tissues.PLoS One. 2013;8(2):e57807. doi: 10.1371/journal.pone.0057807. Epub 2013 Feb 25. PLoS One. 2013. PMID: 23451271 Free PMC article.
-
WNK4 kinase stimulates caveola-mediated endocytosis of TRPV5 amplifying the dynamic range of regulation of the channel by protein kinase C.J Biol Chem. 2010 Feb 26;285(9):6604-11. doi: 10.1074/jbc.M109.056044. Epub 2010 Jan 8. J Biol Chem. 2010. PMID: 20061383 Free PMC article.
-
Emerging roles for WNK kinases in cancer.Cell Mol Life Sci. 2010 Apr;67(8):1265-76. doi: 10.1007/s00018-010-0261-6. Epub 2010 Jan 22. Cell Mol Life Sci. 2010. PMID: 20094755 Free PMC article. Review.
-
Thirty years of the NaCl cotransporter: from cloning to physiology and structure.Am J Physiol Renal Physiol. 2023 Oct 1;325(4):F479-F490. doi: 10.1152/ajprenal.00114.2023. Epub 2023 Aug 10. Am J Physiol Renal Physiol. 2023. PMID: 37560773 Free PMC article. Review.
References
-
- Achard, J. M., D. G. Warnock, S. Disse-Nicodeme, B. Fiquet-Kempf, P. Corvol, A. Fournier, and X. Jeunemaitre. 2003. Familial hyperkalemic hypertension: phenotypic analysis in a large family with the WNK1 deletion mutation. Am. J. Med. 114:495-498. - PubMed
-
- Chomczynski, P., and N. Sacchi. 1987. Single-step method of RNA isolation by acid guanidinium thiocyanate-phenol-chloroform extraction. Anal. Biochem. 162:156-159. - PubMed
-
- Disse-Nicodeme, S., J. M. Achard, J. Potier, M. Delahousse, B. Fiquet-Kempf, N. Stern, A. Blanchard, J. C. Guilbaud, P. Niaudet, D. Chauveau, B. Dussol, Y. Berland, P. Dequiedt, J. L. Ader, M. Paillard, J. P. Grunfeld, A. Fournier, P. Corvol, and X. Jeunemaitre. 2001. Familial hyperkalemic hypertension (Gordon syndrome): evidence for phenotypic variability in a study of 7 families. Adv. Nephrol. Necker Hosp. 31:55-68. - PubMed
-
- Gordon, R. D., S. A. Klemm, T. J. Tunny, and M. Stowasser. 1995. Gordon's syndrome: a sodium-volume-dependent form of hypertension with a genetic basis, p. 2111-2123. In J. H. Laragh and B. M. Brenner (ed.), Hypertension: pathophysiology, diagnosis and management, 2nd ed. Raven Press, New York, N.Y.
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
