Capillary malformation-arteriovenous malformation, a new clinical and genetic disorder caused by RASA1 mutations
- PMID: 14639529
- PMCID: PMC1180390
- DOI: 10.1086/379793
Capillary malformation-arteriovenous malformation, a new clinical and genetic disorder caused by RASA1 mutations
Abstract
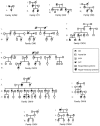
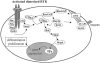
Capillary malformation (CM), or "port-wine stain," is a common cutaneous vascular anomaly that initially appears as a red macular stain that darkens over years. CM also occurs in several combined vascular anomalies that exhibit hypertrophy, such as Sturge-Weber syndrome, Klippel-Trenaunay syndrome, and Parkes Weber syndrome. Occasional familial segregation of CM suggests that there is genetic susceptibility, underscored by the identification of a large locus, CMC1, on chromosome 5q. We used genetic fine mapping with polymorphic markers to reduce the size of the CMC1 locus. A positional candidate gene, RASA1, encoding p120-RasGAP, was screened for mutations in 17 families. Heterozygous inactivating RASA1 mutations were detected in six families manifesting atypical CMs that were multiple, small, round to oval in shape, and pinkish red in color. In addition to CM, either arteriovenous malformation, arteriovenous fistula, or Parkes Weber syndrome was documented in all the families with a mutation. We named this newly identified association caused by RASA1 mutations "CM-AVM," for capillary malformation-arteriovenous malformation. The phenotypic variability can be explained by the involvement of p120-RasGAP in signaling for various growth factor receptors that control proliferation, migration, and survival of several cell types, including vascular endothelial cells.
Figures



Similar articles
-
RASA1 mutations and associated phenotypes in 68 families with capillary malformation-arteriovenous malformation.Hum Mutat. 2013 Dec;34(12):1632-41. doi: 10.1002/humu.22431. Epub 2013 Oct 10. Hum Mutat. 2013. PMID: 24038909
-
RASA1 somatic mutation and variable expressivity in capillary malformation/arteriovenous malformation (CM/AVM) syndrome.Am J Med Genet A. 2016 Jun;170(6):1450-4. doi: 10.1002/ajmg.a.37613. Epub 2016 Mar 11. Am J Med Genet A. 2016. PMID: 26969842
-
Somatic second hit mutation of RASA1 in vascular endothelial cells in capillary malformation-arteriovenous malformation.Eur J Med Genet. 2018 Jan;61(1):11-16. doi: 10.1016/j.ejmg.2017.10.004. Epub 2017 Oct 9. Eur J Med Genet. 2018. PMID: 29024832 Free PMC article.
-
RhoA activation-mediated vascular permeability in capillary malformation-arteriovenous malformation syndrome: a hypothesis.Drug Discov Today. 2021 Aug;26(8):1790-1793. doi: 10.1016/j.drudis.2020.12.012. Epub 2020 Dec 24. Drug Discov Today. 2021. PMID: 33358701 Review.
-
[Neonatal capillary malformation-arteriovenous malformation complicated with acute heart failure: a case report and literature review].Zhonghua Er Ke Za Zhi. 2020 Jul 2;58(7):591-595. doi: 10.3760/cma.j.cn112140-20200312-00221. Zhonghua Er Ke Za Zhi. 2020. PMID: 32605345 Review. Chinese.
Cited by
-
Updates on Sturge-Weber Syndrome.Stroke. 2022 Dec;53(12):3769-3779. doi: 10.1161/STROKEAHA.122.038585. Epub 2022 Oct 20. Stroke. 2022. PMID: 36263782 Free PMC article. Review.
-
Recent progress understanding pathophysiology and genesis of brain AVM-a narrative review.Neurosurg Rev. 2021 Dec;44(6):3165-3175. doi: 10.1007/s10143-021-01526-0. Epub 2021 Apr 10. Neurosurg Rev. 2021. PMID: 33837504 Free PMC article. Review.
-
Cardiac and vascular functions of the zebrafish orthologues of the type I neurofibromatosis gene NFI.Proc Natl Acad Sci U S A. 2009 Dec 29;106(52):22305-10. doi: 10.1073/pnas.0901932106. Epub 2009 Dec 4. Proc Natl Acad Sci U S A. 2009. PMID: 19966217 Free PMC article.
-
Small GTPases and Their Regulators: A Leading Road toward Blood Vessel Development in Zebrafish.Int J Mol Sci. 2022 Apr 30;23(9):4991. doi: 10.3390/ijms23094991. Int J Mol Sci. 2022. PMID: 35563380 Free PMC article. Review.
-
Vascular malformations involving the female pelvis.Semin Intervent Radiol. 2008 Dec;25(4):347-60. doi: 10.1055/s-0028-1102993. Semin Intervent Radiol. 2008. PMID: 21326576 Free PMC article.
References
Electronic-Database Information
-
- NCBI Entrez, http://www.ncbi.nlm.nih.gov/entrez/query.fcgi (for RASA1 cDNA and genomic sequences)
-
- Online Mendelian Inheritance in Man (OMIM), http://www.ncbi.nlm.nih.gov/Omim/ (for CM and CCM) - PubMed
-
- PROSITE, http://us.expasy.org/prosite/ (for PH-domain searches)
References
-
- Ballaun C, Weninger W, Uthman A, Weich H, Tschachler E (1995) Human keratinocytes express the three major splice forms of vascular endothelial growth factor. J Invest Dermatol 104:7–10 - PubMed
-
- Ballester R, Marchuk D, Boguski M, Saulino A, Letcher R, Wigler M, Collins F (1990) The NF1 locus encodes a protein functionally related to mammalian GAP and yeast IRA proteins. Cell 63:851–859 - PubMed
-
- Barsky SH, Rosen S, Geer DE, Noe JM (1980) The nature and evolution of port wine stains: a computer-assisted study. J Invest Dermatol 74:154–157 - PubMed
-
- Bos JL (1989) ras oncogenes in human cancer: a review. Cancer Res 49:4682–2689 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Miscellaneous

