Autosomal dominant pseudohypoparathyroidism type Ib is associated with a heterozygous microdeletion that likely disrupts a putative imprinting control element of GNAS
- PMID: 14561710
- PMCID: PMC213493
- DOI: 10.1172/JCI19159
Autosomal dominant pseudohypoparathyroidism type Ib is associated with a heterozygous microdeletion that likely disrupts a putative imprinting control element of GNAS
Abstract
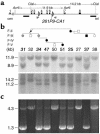
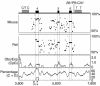
Patients with pseudohypoparathyroidism type Ib (PHP-Ib) have hypocalcemia and hyperphosphatemia due to renal parathyroid hormone (PTH) resistance, but lack physical features of Albright hereditary osteodystrophy. PHP-Ib is thus distinct from PHP-Ia, which is caused by mutations in the GNAS exons encoding the G protein alpha subunit. However, an imprinted autosomal dominant form of PHP-Ib (AD-PHP-Ib) has been mapped to a region of chromosome 20q13.3 containing GNAS. Furthermore, loss of methylation at a differentially methylated region (DMR) of this locus, exon A/B, has been observed thus far in all investigated sporadic PHP-Ib cases and the affected members of multiple AD-PHP-Ib kindreds. We now report that affected members and obligate gene carriers of 12 unrelated AD-PHP-Ib kindreds and four apparently sporadic PHP-Ib patients, but not healthy controls, have a heterozygous approximately 3-kb microdeletion located approximately 220 kb centromeric of GNAS exon A/B. The deleted region, which is flanked by two direct repeats, includes three exons of STX16, the gene encoding syntaxin-16, for which no evidence of imprinting could be found. Affected individuals carrying the microdeletion show loss of exon A/B methylation but no epigenetic abnormalities at other GNAS DMRs. We therefore postulate that this microdeletion disrupts a putative cis-acting element required for methylation at exon A/B, and that this genetic defect underlies the renal PTH resistance in AD-PHP-Ib.
Figures
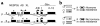


Similar articles
-
A novel STX16 deletion in autosomal dominant pseudohypoparathyroidism type Ib redefines the boundaries of a cis-acting imprinting control element of GNAS.Am J Hum Genet. 2005 May;76(5):804-14. doi: 10.1086/429932. Epub 2005 Mar 30. Am J Hum Genet. 2005. PMID: 15800843 Free PMC article.
-
The GNAS locus and pseudohypoparathyroidism.Adv Exp Med Biol. 2008;626:27-40. doi: 10.1007/978-0-387-77576-0_3. Adv Exp Med Biol. 2008. PMID: 18372789 Review.
-
Lack of Gnas epigenetic changes and pseudohypoparathyroidism type Ib in mice with targeted disruption of syntaxin-16.Endocrinology. 2007 Jun;148(6):2925-35. doi: 10.1210/en.2006-1298. Epub 2007 Feb 22. Endocrinology. 2007. PMID: 17317779
-
GNAS epigenetic defects and pseudohypoparathyroidism: time for a new classification?Horm Metab Res. 2012 Sep;44(10):716-23. doi: 10.1055/s-0032-1314842. Epub 2012 Jun 6. Horm Metab Res. 2012. PMID: 22674477 Review.
-
Phenotypic and molecular genetic aspects of pseudohypoparathyroidism type Ib in a Greek kindred: evidence for enhanced uric acid excretion due to parathyroid hormone resistance.J Clin Endocrinol Metab. 2004 Dec;89(12):5942-7. doi: 10.1210/jc.2004-0249. J Clin Endocrinol Metab. 2004. PMID: 15579741
Cited by
-
Gene Dosage Effects at the Imprinted Gnas Cluster.PLoS One. 2013 Jun 18;8(6):e65639. doi: 10.1371/journal.pone.0065639. Print 2013. PLoS One. 2013. PMID: 23822972 Free PMC article.
-
GNAS Spectrum of Disorders.Curr Osteoporos Rep. 2015 Jun;13(3):146-58. doi: 10.1007/s11914-015-0268-x. Curr Osteoporos Rep. 2015. PMID: 25851935 Free PMC article. Review.
-
A novel de novo 20q13.32-q13.33 deletion in a 2-year-old child with poor growth, feeding difficulties and low bone mass.J Hum Genet. 2015 Jun;60(6):313-7. doi: 10.1038/jhg.2015.22. Epub 2015 Mar 12. J Hum Genet. 2015. PMID: 25761574
-
Human Reproduction and Disturbed Genomic Imprinting.Genes (Basel). 2024 Jan 26;15(2):163. doi: 10.3390/genes15020163. Genes (Basel). 2024. PMID: 38397153 Free PMC article. Review.
-
Identification of the control region for tissue-specific imprinting of the stimulatory G protein alpha-subunit.Proc Natl Acad Sci U S A. 2005 Apr 12;102(15):5513-8. doi: 10.1073/pnas.0408262102. Epub 2005 Apr 5. Proc Natl Acad Sci U S A. 2005. PMID: 15811946 Free PMC article.
References
-
- Levine, M.A. 2002. Pseudohypoparathyroidism. In Principles of bone biology. J.P. Bilezikian, L.G. Raisz, and G.A. Rodan, editors. Academic Press. New York, New York, USA. 1137–1163.
-
- Weinstein LS, Yu S, Warner DR, Liu J. Endocrine manifestations of stimulatory G protein alpha-subunit mutations and the role of genomic imprinting. Endocr. Rev. 2001;22:675–705. - PubMed
-
- Kehlenbach RH, Matthey J, Huttner WB. XLαs is a new type of G protein [erratum 1995, 375:253] Nature. 1994;372:804–809. - PubMed
-
- Pasolli H, Klemke M, Kehlenbach R, Wang Y, Huttner W. Characterization of the extra-large G protein alpha-subunit XLalphas. I. Tissue distribution and subcellular localization. J. Biol. Chem. 2000;275:33622–33632. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

