Functional dissection of eyes absent reveals new modes of regulation within the retinal determination gene network
- PMID: 12917324
- PMCID: PMC180989
- DOI: 10.1128/MCB.23.17.5989-5999.2003
Functional dissection of eyes absent reveals new modes of regulation within the retinal determination gene network
Abstract
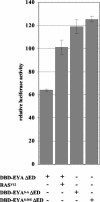
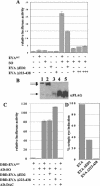
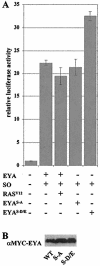
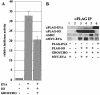
The retinal determination (RD) gene network encodes a group of transcription factors and cofactors necessary for eye development. Transcriptional and posttranslational regulation of RD family members is achieved through interactions within the network and with extracellular signaling pathways, including epidermal growth factor receptor/RAS/mitogen-activated protein kinase (MAPK), transforming growth factor beta/DPP, Wingless, Hedgehog, and Notch. Here we present the results of structure-function analyses that reveal novel aspects of Eyes absent (EYA) function and regulation. We find that the conserved C-terminal EYA domain negatively regulates EYA transactivation potential, and that GROUCHO-SINE OCULIS (SO) interactions provide another mechanism for negative regulation of EYA-SO target genes. We have mapped the transactivation potential of EYA to an internal proline-, serine-, and threonine-rich region that includes the EYA domain 2 (ED2) and two MAPK phosphorylation consensus sites and demonstrate that activation of the RAS/MAPK pathway potentiates transcriptional output of EYA and the EYA-SO complex in certain contexts. Drosophila S2 cell two-hybrid assays were used to describe a novel homotypic interaction that is mediated by EYA's N terminus. Our data suggest that EYA requires homo- and heterotypic interactions and RAS/MAPK signaling responsiveness to ensure context-appropriate RD gene network activity.
Figures


Similar articles
-
Nemo phosphorylates Eyes absent and enhances output from the Eya-Sine oculis transcriptional complex during Drosophila retinal determination.Dev Biol. 2012 May 1;365(1):267-76. doi: 10.1016/j.ydbio.2012.02.030. Epub 2012 Feb 25. Dev Biol. 2012. PMID: 22394486 Free PMC article.
-
Drosophila eyes absent is required for normal cone and pigment cell development.PLoS One. 2014 Jul 24;9(7):e102143. doi: 10.1371/journal.pone.0102143. eCollection 2014. PLoS One. 2014. PMID: 25057928 Free PMC article.
-
Direct control of the proneural gene atonal by retinal determination factors during Drosophila eye development.Dev Biol. 2008 Jan 15;313(2):787-801. doi: 10.1016/j.ydbio.2007.11.017. Epub 2007 Nov 28. Dev Biol. 2008. PMID: 18083159 Free PMC article.
-
Regulation of cancer stem cell properties by SIX1, a member of the PAX-SIX-EYA-DACH network.Adv Cancer Res. 2019;141:1-42. doi: 10.1016/bs.acr.2018.12.001. Epub 2019 Jan 16. Adv Cancer Res. 2019. PMID: 30691681 Review.
-
The six family of homeobox genes in development and cancer.Adv Cancer Res. 2008;101:93-126. doi: 10.1016/S0065-230X(08)00405-3. Adv Cancer Res. 2008. PMID: 19055944 Review.
Cited by
-
Retinal Expression of the Drosophila eyes absent Gene Is Controlled by Several Cooperatively Acting Cis-regulatory Elements.PLoS Genet. 2016 Dec 8;12(12):e1006462. doi: 10.1371/journal.pgen.1006462. eCollection 2016 Dec. PLoS Genet. 2016. PMID: 27930646 Free PMC article.
-
Nemo phosphorylates Eyes absent and enhances output from the Eya-Sine oculis transcriptional complex during Drosophila retinal determination.Dev Biol. 2012 May 1;365(1):267-76. doi: 10.1016/j.ydbio.2012.02.030. Epub 2012 Feb 25. Dev Biol. 2012. PMID: 22394486 Free PMC article.
-
The EYA-SO/SIX complex in development and disease.Pediatr Nephrol. 2013 Jun;28(6):843-54. doi: 10.1007/s00467-012-2246-1. Epub 2012 Jul 19. Pediatr Nephrol. 2013. PMID: 22806561 Free PMC article. Review.
-
Dual transcriptional activities of SIX proteins define their roles in normal and ectopic eye development.Development. 2012 Mar;139(5):991-1000. doi: 10.1242/dev.077255. Development. 2012. PMID: 22318629 Free PMC article.
-
The transcription factor six1 inhibits neuronal and promotes hair cell fate in the developing zebrafish (Danio rerio) inner ear.J Neurosci. 2006 Oct 11;26(41):10438-51. doi: 10.1523/JNEUROSCI.1025-06.2006. J Neurosci. 2006. PMID: 17035528 Free PMC article.
References
-
- Abdelhak, S., V. Kalatzis, R. Heilig, S. Compain, D. Samson, C. Vincent, D. Weil, C. Cruaud, I. Sahly, M. Leibovici, M. Bitner-Glindzicz, M. Francis, D. Lacombe, J. Vigneron, R. Charachon, K. Boven, P. Bedbeder, N. Van Regemorter, J. Weissenbach, and C. Petit. 1997. A human homologue of the Drosophila eyes absent gene underlies branchio-oto-renal (BOR) syndrome and identifies a novel gene family. Nat. Genet. 15:157-164. - PubMed
-
- Bai, J., and D. Montell. 2002. Eyes Absent, a key repressor of polar cell fate during Drosophila oogenesis. Development 129:5377-5388. - PubMed
-
- Baonza, A., and M. Freeman. 2002. Control of Drosophila eye specification by Wingless signalling. Development 129:5313-5322. - PubMed
-
- Bonini, N. M., Q. T. Bui, G. L. Gray-Board, and J. M. Warrick. 1997. The Drosophila eyes absent gene directs ectopic eye formation in a pathway conserved between flies and vertebrates. Development 124:4819-4826. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous
