Mutational mechanisms of Williams-Beuren syndrome deletions
- PMID: 12796854
- PMCID: PMC1180575
- DOI: 10.1086/376565
Mutational mechanisms of Williams-Beuren syndrome deletions
Abstract
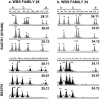
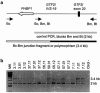
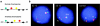
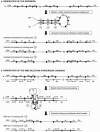
Williams-Beuren syndrome (WBS) is a segmental aneusomy syndrome that results from a heterozygous deletion of contiguous genes at 7q11.23. Three large region-specific low-copy repeat elements (LCRs), composed of different blocks (A, B, and C), flank the WBS deletion interval and are thought to predispose to misalignment and unequal crossing-over, causing the deletions. In this study, we have determined the exact deletion size and LCR copy number in 74 patients with WBS, as well as precisely defined deletion breakpoints in 30 of them, using LCR-specific nucleotide differences. Most patients (95%) exhibit a 1.55-Mb deletion caused by recombination between centromeric and medial block B copies, which share approximately 99.6% sequence identity along 105-143 kb. In these cases, deletion breakpoints were mapped at several sites within the recombinant block B, with a cluster (>27%) occurring at a 12 kb region within the GTF2I/GTF2IP1 gene. Almost one-third (28%) of the transmitting progenitors were found to be heterozygous for an inversion between centromeric and telomeric LCRs. All deletion breakpoints in the patients with the inversion occurred in the distal 38-kb block B region only present in the telomeric and medial copies. Finally, only four patients (5%) displayed a larger deletion ( approximately 1.84 Mb) caused by recombination between centromeric and medial block A copies. We propose models for the specific pairing and precise aberrant recombination leading to each of the different germline rearrangements that occur in this region, including inversions and deletions associated with WBS. Chromosomal instability at 7q11.23 is directly related to the genomic structure of the region.
Figures
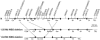



Similar articles
-
A duplicated gene in the breakpoint regions of the 7q11.23 Williams-Beuren syndrome deletion encodes the initiator binding protein TFII-I and BAP-135, a phosphorylation target of BTK.Hum Mol Genet. 1998 Mar;7(3):325-34. doi: 10.1093/hmg/7.3.325. Hum Mol Genet. 1998. PMID: 9466987
-
Copy number variation at the 7q11.23 segmental duplications is a susceptibility factor for the Williams-Beuren syndrome deletion.Genome Res. 2008 May;18(5):683-94. doi: 10.1101/gr.073197.107. Epub 2008 Feb 21. Genome Res. 2008. PMID: 18292220 Free PMC article.
-
Fine-scale comparative mapping of the human 7q11.23 region and the orthologous region on mouse chromosome 5G: the low-copy repeats that flank the Williams-Beuren syndrome deletion arose at breakpoint sites of an evolutionary inversion(s).Genomics. 2000 Oct 1;69(1):1-13. doi: 10.1006/geno.2000.6312. Genomics. 2000. PMID: 11013070
-
The genomic basis of the Williams-Beuren syndrome.Cell Mol Life Sci. 2009 Apr;66(7):1178-97. doi: 10.1007/s00018-008-8401-y. Cell Mol Life Sci. 2009. PMID: 19039520 Free PMC article. Review.
-
Williams-Beuren syndrome: genes and mechanisms.Hum Mol Genet. 1999;8(10):1947-54. doi: 10.1093/hmg/8.10.1947. Hum Mol Genet. 1999. PMID: 10469848 Review.
Cited by
-
Low copy repeats mediate distal chromosome 22q11.2 deletions: sequence analysis predicts breakpoint mechanisms.Genome Res. 2007 Apr;17(4):482-91. doi: 10.1101/gr.5986507. Epub 2007 Mar 9. Genome Res. 2007. PMID: 17351135 Free PMC article.
-
Inversion of the Williams syndrome region is a common polymorphism found more frequently in parents of children with Williams syndrome.Am J Med Genet C Semin Med Genet. 2010 May 15;154C(2):220-8. doi: 10.1002/ajmg.c.30258. Am J Med Genet C Semin Med Genet. 2010. PMID: 20425783 Free PMC article.
-
Two high throughput technologies to detect segmental aneuploidies identify new Williams-Beuren syndrome patients with atypical deletions.J Med Genet. 2006 Mar;43(3):266-73. doi: 10.1136/jmg.2005.034009. Epub 2005 Jul 1. J Med Genet. 2006. PMID: 15994861 Free PMC article.
-
Leftward lateralization of auditory cortex underlies holistic sound perception in Williams syndrome.PLoS One. 2010 Aug 23;5(8):e12326. doi: 10.1371/journal.pone.0012326. PLoS One. 2010. PMID: 20808792 Free PMC article.
-
A human genome structural variation sequencing resource reveals insights into mutational mechanisms.Cell. 2010 Nov 24;143(5):837-47. doi: 10.1016/j.cell.2010.10.027. Cell. 2010. PMID: 21111241 Free PMC article.
References
Electronic-Database Information
-
- BLAST search engine, http://www.ncbi.nlm.nih.gov/BLAST/
-
- DNAsp 3.51, http://www.ub.es/dnasp/
-
- GenBank, http://www.ncbi.nih.gov/GenBank/ (for BACs CTA-269p13 [accession number AC005080], RP11-396k3 [accession number AC006995], RP11-450o3 [accession number AC105418, RP11-483g21] [accession number AC00416], 239c10 [accession number AC004883], RP4-771p04 [accession number AC083884], RP11-813J7 [accession number AC005098], CTA-350l10 [accession number AC124781], RP11-729p19 [accession number AC027219], and RP11-219m8 [accession number AC124781]; prohibitin-related ESTs from fetal brain [accession number AA076811]; and stomach cancer [accession number AW814764])
-
- Genome Database, http://gdbwww.gdb.org/ for primer information on HSB055XE5 [ID number 609780], D7S489 [ID number 188049], D7S653 [ID number 199574], D7S672 [ID number 199800], D7S1816 [ID number 684408], D7S2518 [ID number 612411], and D7S1870 [ID number 377150])
References
-
- Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410 - PubMed
-
- Amos-Landgraf JM, Ji Y, Gottlieb W, Depinet T, Wandstrat AE, Cassidy SB, Driscoll DJ, Rogan PK, Schwartz S, Nicholls RD (1999) Chromosome breakage in the Prader-Willi and Angelman syndromes involves recombination between large, transcribed repeats at proximal and distal breakpoints. Am J Hum Genet 65:370–386 - PMC - PubMed
-
- Batzer MA, Deininger PL (2002) Alu repeats and human genomic diversity. Nat Rev Genet 3:370–379 - PubMed
-
- Baumer A, Dutly F, Balmer D, Riegel M, Tükel T, Krajewska-Walasek M, Schinzel AA (1998) High level of unequal meiotic crossovers at the origin of the 22q11.2 and 7q11.23 deletions. Hum Mol Genet 7:887–984 - PubMed
-
- Bellugi U, Lichtenberger L, Jones W, Lai Z, St George M (2000) The neurocognitive profile of Williams Syndrome: a complex pattern of strengths and weaknesses. J Cogn Neurosci Suppl 12:7–29 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

