Adenosine deaminase-related growth factors stimulate cell proliferation in Drosophila by depleting extracellular adenosine
- PMID: 11904370
- PMCID: PMC123660
- DOI: 10.1073/pnas.062059699
Adenosine deaminase-related growth factors stimulate cell proliferation in Drosophila by depleting extracellular adenosine
Abstract
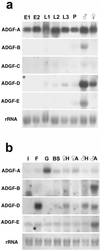
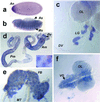
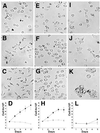
We describe a protein family in Drosophila containing six adenosine deaminase-related growth factors (ADGFs), which are homologous to a mitogenic growth factor discovered in conditioned medium from cells of a different fly species, Sarcophaga. Closely related proteins have been identified in other animals, and a human homolog is implicated in the genetic disease Cat-Eye Syndrome. The two most abundantly expressed ADGFs in Drosophila larvae are ADGF-A, which is strongly expressed in the gut and lymph glands, and ADGF-D, which is mainly expressed in the fat body and brain. Recombinant ADGF-A and ADGF-D are active adenosine deaminases (ADAs), and they cause polarization and serum-independent proliferation of imaginal disk and embryonic cells in vitro. The enzymatic activity of these proteins is required for their mitogenic function, making them unique among growth factors. A culture medium prepared without adenosine, or depleted of adenosine by using bovine ADA, also stimulates proliferation of imaginal disk cells, and addition of adenosine to this medium inhibits proliferation. Thus ADGFs secreted in vivo may control tissue growth by modulating the level of extracellular adenosine.
Figures


Similar articles
-
Phylogenetic analysis reveals a novel protein family closely related to adenosine deaminase.J Mol Evol. 2005 Dec;61(6):776-94. doi: 10.1007/s00239-005-0046-y. Epub 2005 Oct 20. J Mol Evol. 2005. PMID: 16245011
-
Mollusk-derived growth factor and the new subfamily of adenosine deaminase-related growth factors.Curr Pharm Des. 2004;10(31):3893-900. doi: 10.2174/1381612043382594. Curr Pharm Des. 2004. PMID: 15579078 Review.
-
The AMP deaminase of the mollusk Helix pomatia is an unexpected member of the adenosine deaminase-related growth factor (ADGF) family.PLoS One. 2023 Jul 20;18(7):e0286435. doi: 10.1371/journal.pone.0286435. eCollection 2023. PLoS One. 2023. PMID: 37471401 Free PMC article.
-
Secreted Bacterial Adenosine Deaminase Is an Evolutionary Precursor of Adenosine Deaminase Growth Factor.Mol Biol Evol. 2018 Dec 1;35(12):2851-2861. doi: 10.1093/molbev/msy193. Mol Biol Evol. 2018. PMID: 30325443
-
The DRE/DREF transcriptional regulatory system: a master key for cell proliferation.Biochim Biophys Acta. 2008 Feb;1779(2):81-9. doi: 10.1016/j.bbagrm.2007.11.011. Epub 2007 Dec 4. Biochim Biophys Acta. 2008. PMID: 18155677 Review.
Cited by
-
Targeted genetics in Drosophila cell lines: Inserting single transgenes in vitro.Fly (Austin). 2016 Jul 2;10(3):134-41. doi: 10.1080/19336934.2016.1191716. Epub 2016 Jun 3. Fly (Austin). 2016. PMID: 27261098 Free PMC article.
-
Genomic disorders on 22q11.Am J Hum Genet. 2002 May;70(5):1077-88. doi: 10.1086/340363. Epub 2002 Mar 29. Am J Hum Genet. 2002. PMID: 11925570 Free PMC article. Review.
-
Structure of Arabidopsis thaliana N6-methyl-AMP deaminase ADAL with bound GMP and IMP and implications for N6-methyl-AMP recognition and processing.RNA Biol. 2019 Oct;16(10):1504-1512. doi: 10.1080/15476286.2019.1642712. Epub 2019 Jul 18. RNA Biol. 2019. PMID: 31318636 Free PMC article.
-
Phylogenetic analysis reveals a novel protein family closely related to adenosine deaminase.J Mol Evol. 2005 Dec;61(6):776-94. doi: 10.1007/s00239-005-0046-y. Epub 2005 Oct 20. J Mol Evol. 2005. PMID: 16245011
-
Characterization and Expression Analysis of Mollusk-like Growth Factor: A Secreted Protein Involved in Pacific Abalone Embryonic and Larval Development.Biology (Basel). 2022 Oct 1;11(10):1445. doi: 10.3390/biology11101445. Biology (Basel). 2022. PMID: 36290349 Free PMC article.
References
-
- Bryant P J, Simpson P. Q Rev Biol. 1984;59:387–415. - PubMed
-
- Diaz-Benjumea F J, Cohen B, Cohen S M. Nature (London) 1994;372:175–179. - PubMed
-
- Cross M, Dexter T M. Cell. 1991;64:271–280. - PubMed
-
- Kawamura K, Shibata T, Saget O, Peel D, Bryant P J. Development. 1999;126:211–219. - PubMed
-
- Wintrobe M M. Clinical Hematology. Philadelphia: Lea & Febiger; 1956. pp. 1–1185.
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

