The Pseudomonas aeruginosa devB/SOL homolog, pgl, is a member of the hex regulon and encodes 6-phosphogluconolactonase
- PMID: 10869070
- PMCID: PMC94577
- DOI: 10.1128/JB.182.14.3934-3941.2000
The Pseudomonas aeruginosa devB/SOL homolog, pgl, is a member of the hex regulon and encodes 6-phosphogluconolactonase
Abstract
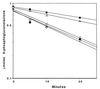
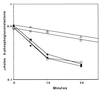
A cyclic version of the Entner-Doudoroff pathway is used by Pseudomonas aeruginosa to metabolize carbohydrates. Genes encoding the enzymes that catabolize intracellular glucose to pyruvate and glyceraldehyde 3-phosphate are coordinately regulated, clustered at 39 min on the chromosome, and collectively form the hex regulon. Within the hex cluster is an open reading frame (ORF) with homology to the devB/SOL family of unidentified proteins. This ORF encodes a protein of either 243 or 238 amino acids; it overlaps the 5' end of zwf (encodes glucose-6-phosphate dehydrogenase) and is followed immediately by eda (encodes the Entner-Doudoroff aldolase). The devB/SOL homolog was inactivated in P. aeruginosa PAO1 by recombination with a suicide plasmid containing an interrupted copy of the gene, creating mutant strain PAO8029. PAO8029 grows at 9% of the wild-type rate using mannitol as the carbon source and at 50% of the wild-type rate using gluconate as the carbon source. Cell extracts of PAO8029 were specifically deficient in 6-phosphogluconolactonase (Pgl) activity. The cloned devB/SOL homolog complemented PAO8029 to restore normal growth on mannitol and gluconate and restored Pgl activity. Hence, we have identified this gene as pgl and propose that the devB/SOL family members encode 6-phosphogluconolactonases. Interestingly, three eukaryotic glucose-6-phosphate dehydrogenase (G6PDH) isozymes, from human, rabbit, and Plasmodium falciparum, contain Pgl domains, suggesting that the sequential reactions of G6PDH and Pgl are incorporated in a single protein. 6-Phosphogluconolactonase activity is induced in P. aeruginosa PAO1 by growth on mannitol and repressed by growth on succinate, and it is expressed constitutively in P. aeruginosa PAO8026 (hexR). Taken together, these results establish that Pgl is an essential enzyme of the cyclic Entner-Doudoroff pathway encoded by pgl, a structural gene of the hex regulon.
Figures
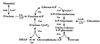




Similar articles
-
Analysis of the zwf-pgl-eda-operon in Pseudomonas putida strains H and KT2440.FEMS Microbiol Lett. 2002 Sep 24;215(1):89-95. doi: 10.1111/j.1574-6968.2002.tb11375.x. FEMS Microbiol Lett. 2002. PMID: 12393206
-
Characterization of the Entner-Doudoroff pathway in Pseudomonas aeruginosa catheter-associated urinary tract infections.J Bacteriol. 2024 Jan 25;206(1):e0036123. doi: 10.1128/jb.00361-23. Epub 2023 Dec 4. J Bacteriol. 2024. PMID: 38047680 Free PMC article.
-
Fructose 1,6-bisphosphate aldolase activity is essential for synthesis of alginate from glucose by Pseudomonas aeruginosa.J Bacteriol. 1985 Jan;161(1):458-60. doi: 10.1128/jb.161.1.458-460.1985. J Bacteriol. 1985. PMID: 3918010 Free PMC article.
-
Two genes for carbohydrate catabolism are divergently transcribed from a region of DNA containing the hexC locus in Pseudomonas aeruginosa PAO1.J Bacteriol. 1994 Aug;176(15):4700-9. doi: 10.1128/jb.176.15.4700-4709.1994. J Bacteriol. 1994. PMID: 8045900 Free PMC article.
-
6-Phosphogluconate dehydratase deficiency in pleiotropic carbohydrate-negative mutant strains of Pseudomonas aeruginosa.J Bacteriol. 1975 Mar;121(3):942-9. doi: 10.1128/jb.121.3.942-949.1975. J Bacteriol. 1975. PMID: 163817 Free PMC article.
Cited by
-
Suppressing posttranslational gluconoylation of heterologous proteins by metabolic engineering of Escherichia coli.Appl Environ Microbiol. 2008 Feb;74(4):950-8. doi: 10.1128/AEM.01790-07. Epub 2007 Dec 14. Appl Environ Microbiol. 2008. PMID: 18083862 Free PMC article.
-
Vibrio fischeri siderophore production drives competitive exclusion during dual-species growth.Mol Microbiol. 2020 Aug;114(2):244-261. doi: 10.1111/mmi.14509. Epub 2020 May 8. Mol Microbiol. 2020. PMID: 32259318 Free PMC article.
-
Engineering of primary carbon metabolism for improved antibiotic production in Streptomyces lividans.Appl Environ Microbiol. 2002 Oct;68(10):4731-9. doi: 10.1128/AEM.68.10.4731-4739.2002. Appl Environ Microbiol. 2002. PMID: 12324314 Free PMC article.
-
Comparative Genomic Analysis of a Methylorubrum rhodesianum MB200 Isolated from Biogas Digesters Provided New Insights into the Carbon Metabolism of Methylotrophic Bacteria.Int J Mol Sci. 2023 Apr 19;24(8):7521. doi: 10.3390/ijms24087521. Int J Mol Sci. 2023. PMID: 37108681 Free PMC article.
-
Identification of the Escherichia coli K-12 ybhE gene as pgl, encoding 6-phosphogluconolactonase.J Bacteriol. 2004 Dec;186(24):8248-53. doi: 10.1128/JB.186.24.8248-8253.2004. J Bacteriol. 2004. PMID: 15576773 Free PMC article.
References
-
- Altschul S F, Gish W, Miller W, Myers E W, Lipman D J. Basic local alignment search tool. J Mol Biol. 1990;215:403–410. - PubMed
-
- Ausubel F M, Brent R, Kingston R E, et al. Current protocols in molecular biology. New York, N.Y: John Wiley & Sons; 1989.
-
- Bauer H P, Srihari T, Jochims J C, Hofer H W. 6-Phosphogluconolactonase: purification, properties and activities in various tissues. Eur J Biochem. 1983;133:163–168. - PubMed
-
- Beutler E, Kuhl W, Gelbart T. Blood cell phosphogluconolactonase: assay and properties. Br J Haematol. 1986;62:577–586. - PubMed
-
- Bradford M M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem. 1976;72:248–254. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources

