cis and trans factors affecting Mos1 mariner evolution and transposition in vitro, and its potential for functional genomics
- PMID: 10637331
- PMCID: PMC102556
- DOI: 10.1093/nar/28.3.784
cis and trans factors affecting Mos1 mariner evolution and transposition in vitro, and its potential for functional genomics
Abstract
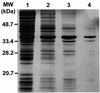
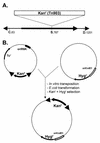
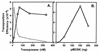
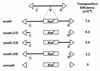
Mos1 and other mariner / Tc1 transposons move horizon-tally during evolution, and when transplanted into heterologous species can transpose in organisms ranging from prokaryotes to protozoans and vertebrates. To further develop the Drosophila Mos1 mariner system as a genetic tool and to probe mechanisms affecting the regulation of transposition activity, we developed an in vitro system for Mos1 transposition using purified transposase and selectable Mos1 derivatives. Transposition frequencies of nearly 10(-3)/target DNA molecule were obtained, and insertions occurred at TA dinucleotides with little other sequence specificity. Mos1 elements containing only the 28 bp terminal inverted repeats were inactive in vitro, while elements containing a few additional internal bases were fully active, establishing the minimal cis -acting requirements for transposition. With increasing transposase the transposition frequency increased to a plateau value, in contrast to the predictions of the protein over-expression inhibition model and to that found recently with a reconstructed Himar1 transposase. This difference between the 'natural' Mos1 and 'reconstructed' Himar1 transposases suggests an evolutionary path for down-regulation of mariner transposition following its introduction into a naïve population. The establishment of the cis and trans requirements for optimal mariner transposition in vitro provides key data for the creation of vectors for in vitro mutagenesis, and will facilitate the development of in vivo systems for mariner transposition.
Figures

Similar articles
-
Regulation of mariner transposition: the peculiar case of Mos1.PLoS One. 2012;7(8):e43365. doi: 10.1371/journal.pone.0043365. Epub 2012 Aug 14. PLoS One. 2012. PMID: 22905263 Free PMC article.
-
DNA-binding activity and subunit interaction of the mariner transposase.Nucleic Acids Res. 2001 Sep 1;29(17):3566-75. doi: 10.1093/nar/29.17.3566. Nucleic Acids Res. 2001. PMID: 11522826 Free PMC article.
-
Physical properties of DNA components affecting the transposition efficiency of the mariner Mos1 element.Mol Genet Genomics. 2009 Nov;282(5):531-46. doi: 10.1007/s00438-009-0484-0. Epub 2009 Sep 23. Mol Genet Genomics. 2009. PMID: 19774400
-
Assembly of the Tc1 and mariner transposition initiation complexes depends on the origins of their transposase DNA binding domains.Genetica. 2007 Jun;130(2):105-20. doi: 10.1007/s10709-006-0025-2. Epub 2006 Aug 16. Genetica. 2007. PMID: 16912840 Review.
-
Research progress of Tc1/Mariner superfamily.Yi Chuan. 2017 Jan 20;39(1):1-13. doi: 10.16288/j.yczz.16-160. Yi Chuan. 2017. PMID: 28115300 Review.
Cited by
-
Transposition of cyanobacterium insertion element ISY100 in Escherichia coli.J Bacteriol. 2002 Sep;184(18):5104-12. doi: 10.1128/JB.184.18.5104-5112.2002. J Bacteriol. 2002. PMID: 12193627 Free PMC article.
-
Unexpected stability of mariner transgenes in Drosophila.Genetics. 2002 Feb;160(2):527-35. doi: 10.1093/genetics/160.2.527. Genetics. 2002. PMID: 11861559 Free PMC article.
-
Biochemical characterization and comparison of two closely related active mariner transposases.Biochemistry. 2014 Feb 4;53(4):682-9. doi: 10.1021/bi401193w. Epub 2014 Jan 21. Biochemistry. 2014. PMID: 24404958 Free PMC article.
-
DNA transposon-based gene vehicles - scenes from an evolutionary drive.J Biomed Sci. 2013 Dec 9;20(1):92. doi: 10.1186/1423-0127-20-92. J Biomed Sci. 2013. PMID: 24320156 Free PMC article. Review.
-
A transposon-based tool for transformation and mutagenesis in trypanosomatid protozoa.Methods Mol Biol. 2015;1201:235-45. doi: 10.1007/978-1-4939-1438-8_14. Methods Mol Biol. 2015. PMID: 25388118 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

