Genetic Structure of Hereditary Forms of Diabetes Mellitus in Russia
- PMID: 39859454
- PMCID: PMC11766241
- DOI: 10.3390/ijms26020740
Genetic Structure of Hereditary Forms of Diabetes Mellitus in Russia
Abstract
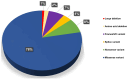
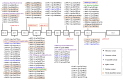
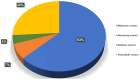
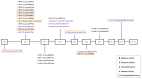
Analyzing the genetic architecture of hereditary forms of diabetes in different populations is a critical step toward optimizing diagnostic and preventive algorithms. This requires consideration of regional and population-specific characteristics, including the spectrum and frequency of pathogenic variants in targeted genes. As part of this study, we used a custom-designed NGS panel to screen for mutations in 28 genes associated with the pathogenesis of hereditary diabetes mellitus in 506 unrelated patients from Russia. The study identified 180 pathogenic or likely pathogenic variants across 13 genes (GCK, HNF1A, HNF1B, HNF4A, ABCC8, INS, INSR, KCNJ11, PAX4, PDX1, ZFP57, BLK, WFS1), representing 46.44% of the analyzed cohort (235 individuals). The glucokinase gene (GCK) had the highest number of identified variants, with 111 variants detected in 161 patients, 20 of which were identified for the first time. In the tissue-specific transcription factor genes HNF1A, HNF4A, and HNF1B, 34 variants were found in 38 patients, including 13 that were previously unreported. Seventeen variants were identified in the ABCC8 gene, which encodes the ATP-binding cassette transporter 8 of subfamily C, each found in a different patient; four of these were novel discoveries. Nine pathogenic or likely pathogenic variants were identified in the insulin gene (INS) and its receptor gene (INSR), including four previously unreported variants. Additionally, we identified 10 previously unreported variants in six other genes among 11 patients. Variants in the genes GCK, HNF1A, HNF1B, HNF4A, ABCC8, INS, and INSR were the main contributors to the genetic pathogenesis of hereditary diabetes mellitus in the Russian cohort. These findings enhance our understanding of the molecular mechanisms underlying the disease and provide a solid basis for future studies aimed at improving diagnostic accuracy and advancing personalized therapeutic strategies. This knowledge provides a foundation for developing region-specific genetic testing algorithms and personalized therapeutic strategies, which are critical for future initiatives in precision medicine.
Keywords: diabetes mellitus; maturity onset diabetes of the young; neonatal diabetes.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
Similar articles
-
Screening of Mutations in Maturity-onset Diabetes of the Young-related Genes and RFX6 in Children with Autoantibody-negative Type 1 Diabetes Mellitus.J Clin Res Pediatr Endocrinol. 2024 May 31;16(2):137-145. doi: 10.4274/jcrpe.galenos.2023.2023-5-10. Epub 2023 Dec 6. J Clin Res Pediatr Endocrinol. 2024. PMID: 38054414 Free PMC article.
-
Defining the optimum strategy for identifying adults and children with coeliac disease: systematic review and economic modelling.Health Technol Assess. 2022 Oct;26(44):1-310. doi: 10.3310/ZUCE8371. Health Technol Assess. 2022. PMID: 36321689 Free PMC article.
-
Our Experiences and Learnings in Diagnosing MODY from Non-Institutional-Based Diabetes Care Clinics.Indian J Endocrinol Metab. 2024 Sep-Oct;28(5):480-487. doi: 10.4103/ijem.ijem_361_23. Epub 2024 Nov 8. Indian J Endocrinol Metab. 2024. PMID: 39676784 Free PMC article.
-
School-based education programmes for the prevention of unintentional injuries in children and young people.Cochrane Database Syst Rev. 2016 Dec 27;12(12):CD010246. doi: 10.1002/14651858.CD010246.pub2. Cochrane Database Syst Rev. 2016. PMID: 28026877 Free PMC article. Review.
-
Interventions to increase COVID-19 vaccine uptake: a scoping review.Cochrane Database Syst Rev. 2022 Aug 3;8(8):CD015270. doi: 10.1002/14651858.CD015270. Cochrane Database Syst Rev. 2022. PMID: 35920693 Free PMC article.
References
-
- Tobias D.K., Merino J., Ahmad A., Aiken C., Benham J.L., Bodhini D., Clark A.L., Colclough K., Corcoy R., Cromer S.J., et al. Second International Consensus Report on Gaps and Opportunities for the Clinical Translation of Precision Diabetes Medicine. Nat. Med. 2023;29:2438–2457. doi: 10.1038/s41591-023-02502-5. - DOI - PMC - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

