The role of exopolysaccharides Psl and Pel in resistance of Pseudomonas aeruginosa to the oxidative stressors sodium hypochlorite and hydrogen peroxide
- PMID: 39194290
- PMCID: PMC11448232
- DOI: 10.1128/spectrum.00922-24
The role of exopolysaccharides Psl and Pel in resistance of Pseudomonas aeruginosa to the oxidative stressors sodium hypochlorite and hydrogen peroxide
Abstract
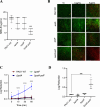
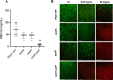
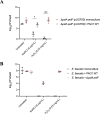
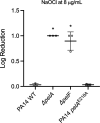
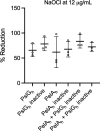
Pseudomonas aeruginosa is well-known for its antimicrobial resistance and the ability to survive in harsh environmental conditions due to an abundance of resistance mechanisms, including the formation of biofilms and the production of exopolysaccharides. Exopolysaccharides are among the major components of the extracellular matrix in biofilms and aggregates of P. aeruginosa. Although their contribution to antibiotic resistance has been previously shown, their roles in resistance to oxidative stressors remain largely elusive. Here, we studied the function of the exopolysaccharides Psl and Pel in the resistance of P. aeruginosa to the commonly used disinfectants and strong oxidizing agents NaOCl and H2O2. We observed that the simultaneous inactivation of Psl and Pel in P. aeruginosa PAO1 mutant strain ∆pslA pelF resulted in a significant increase in susceptibility to both NaOCl and H2O2. Further analyses revealed that Pel is more important for oxidative stress resistance in P. aeruginosa and that the form of Pel (i.e., cell-associated or cell-free) did not affect NaOCl susceptibility. Additionally, we show that Psl/Pel-negative strains are protected against oxidative stress in co-culture biofilms with P. aeruginosa PAO1 WT. Taken together, our results demonstrate that the EPS matrix and, more specifically, Pel exhibit protective functions against oxidative stressors such as NaOCl and H2O2 in P. aeruginosa.
Importance: Biofilms are microbial communities of cells embedded in a self-produced polymeric matrix composed of polysaccharides, proteins, lipids, and extracellular DNA. Biofilm bacteria have been shown to possess unique characteristics, including increased stress resistance and higher antimicrobial tolerance, leading to failures in bacterial eradication during chronic infections or in technical settings, including drinking and wastewater industries. Previous studies have shown that in addition to conferring structure and stability to biofilms, the polysaccharides Psl and Pel are also involved in antibiotic resistance. This work provides evidence that these biofilm matrix components also contribute to the resistance of Pseudomonas aeruginosa to oxidative stressors including the widely used disinfectant NaOCl. Understanding the mechanisms by which bacteria escape antimicrobial agents, including strong oxidants, is urgently needed in the fight against antimicrobial resistance and will help in developing new strategies to eliminate resistant strains in any environmental, industrial, and clinical setting.
Keywords: Pseudomonas aeruginosa; biofilms; exopolysaccharides; hydrogen peroxide; oxidative stress; reactive chlorine species; sodium hypochlorite.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

Similar articles
-
Role of Pel and Psl polysaccharides in the response of Pseudomonas aeruginosa to environmental challenges: oxidative stress agents (UVA, H2O2, sodium hypochlorite) and its competitor Staphylococcus aureus.Microbiology (Reading). 2023 Feb;169(2):001301. doi: 10.1099/mic.0.001301. Microbiology (Reading). 2023. PMID: 36757866 Free PMC article.
-
The Versatile Pseudomonas aeruginosa Biofilm Matrix Protein CdrA Promotes Aggregation through Different Extracellular Exopolysaccharide Interactions.J Bacteriol. 2020 Sep 8;202(19):e00216-20. doi: 10.1128/JB.00216-20. Print 2020 Sep 8. J Bacteriol. 2020. PMID: 32661078 Free PMC article.
-
Genetic determinants of increased sodium hypochlorite and ciprofloxacin susceptibility in Pseudomonas aeruginosa PA14 biofilms.Biofouling. 2024 Oct;40(9):563-579. doi: 10.1080/08927014.2024.2395378. Epub 2024 Aug 27. Biofouling. 2024. PMID: 39189148
-
Recent perspectives on the molecular basis of biofilm formation by Pseudomonas aeruginosa and approaches for treatment and biofilm dispersal.Folia Microbiol (Praha). 2018 Jul;63(4):413-432. doi: 10.1007/s12223-018-0585-4. Epub 2018 Jan 19. Folia Microbiol (Praha). 2018. PMID: 29352409 Review.
-
Pseudomonas aeruginosa biofilm exopolysaccharides: assembly, function, and degradation.FEMS Microbiol Rev. 2023 Nov 1;47(6):fuad060. doi: 10.1093/femsre/fuad060. FEMS Microbiol Rev. 2023. PMID: 37884397 Free PMC article. Review.
References
-
- World Health Organization . 2017. WHO publishes list of bacteria for which new antibiotics are urgently needed. Available from: https://www.who.int/news/item/27-02-2017-who-publishes-list-of-bacteria-.... Retrieved 5 May 2021.
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

