Myxobacterial Genomics and Post-Genomics: A Review of Genome Biology, Genome Sequences and Related 'Omics Studies
- PMID: 34683464
- PMCID: PMC8538405
- DOI: 10.3390/microorganisms9102143
Myxobacterial Genomics and Post-Genomics: A Review of Genome Biology, Genome Sequences and Related 'Omics Studies
Abstract
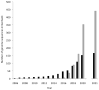
Myxobacteria are fascinating and complex microbes. They prey upon other members of the soil microbiome by secreting antimicrobial proteins and metabolites, and will undergo multicellular development if starved. The genome sequence of the model myxobacterium Myxococcus xanthus DK1622 was published in 2006 and 15 years later, 163 myxobacterial genome sequences have now been made public. This explosion in genomic data has enabled comparative genomics analyses to be performed across the taxon, providing important insights into myxobacterial gene conservation and evolution. The availability of myxobacterial genome sequences has allowed system-wide functional genomic investigations into entire classes of genes. It has also enabled post-genomic technologies to be applied to myxobacteria, including transcriptome analyses (microarrays and RNA-seq), proteome studies (gel-based and gel-free), investigations into protein-DNA interactions (ChIP-seq) and metabolism. Here, we review myxobacterial genome sequencing, and summarise the insights into myxobacterial biology that have emerged as a result. We also outline the application of functional genomics and post-genomic approaches in myxobacterial research, highlighting important findings to emerge from seminal studies. The review also provides a comprehensive guide to the genomic datasets available in mid-2021 for myxobacteria (including 24 genomes that we have sequenced and which are described here for the first time).
Keywords: comparative genomics; functional genomics; genome evolution; genome organisation; pan-genome; proteomics; taxonomy; transcriptomics.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

Similar articles
-
Comparative Genomics and Pan-Genomics of the Myxococcaceae, including a Description of Five Novel Species: Myxococcus eversor sp. nov., Myxococcus llanfairpwllgwyngyllgogerychwyrndrobwllllantysiliogogogochensis sp. nov., Myxococcus vastator sp. nov., Pyxidicoccus caerfyrddinensis sp. nov., and Pyxidicoccus trucidator sp. nov.Genome Biol Evol. 2020 Dec 6;12(12):2289-2302. doi: 10.1093/gbe/evaa212. Genome Biol Evol. 2020. PMID: 33022031 Free PMC article.
-
A survey of non-coding RNAs in the social and predatory myxobacterium Myxococcus xanthus DK1622.Mol Omics. 2020 Oct 12;16(5):492-502. doi: 10.1039/d0mo00068j. Mol Omics. 2020. PMID: 32780046
-
A Genomic Survey of Signalling in the Myxococcaceae.Microorganisms. 2020 Nov 6;8(11):1739. doi: 10.3390/microorganisms8111739. Microorganisms. 2020. PMID: 33171896 Free PMC article.
-
Investigation of cytochromes P450 in myxobacteria: excavation of cytochromes P450 from the genome of Sorangium cellulosum So ce56.FEBS Lett. 2011 Jun 6;585(11):1506-13. doi: 10.1016/j.febslet.2011.04.035. Epub 2011 Apr 22. FEBS Lett. 2011. PMID: 21521637 Review.
-
In depth natural product discovery - Myxobacterial strains that provided multiple secondary metabolites.Biotechnol Adv. 2020 Mar-Apr;39:107480. doi: 10.1016/j.biotechadv.2019.107480. Epub 2019 Nov 7. Biotechnol Adv. 2020. PMID: 31707075 Review.
Cited by
-
Identification of secondary metabolites containing a diketopiperazine core in extracts from myxobacterial strains with growth inhibition activity against a range of prey species.Access Microbiol. 2023 Oct 5;5(10):000629.v4. doi: 10.1099/acmi.0.000629.v4. eCollection 2023. Access Microbiol. 2023. PMID: 37970077 Free PMC article.
-
Phylogenetic Revisit to a Review on Predatory Bacteria.Microorganisms. 2023 Jun 27;11(7):1673. doi: 10.3390/microorganisms11071673. Microorganisms. 2023. PMID: 37512846 Free PMC article. Review.
-
MyxoPortal: a database of myxobacterial genomic features.Database (Oxford). 2024 Jul 2;2024:baae056. doi: 10.1093/database/baae056. Database (Oxford). 2024. PMID: 38958433 Free PMC article.
-
Predatory Strategies of Myxococcus xanthus: Prey Susceptibility to OMVs and Moonlighting Enzymes.Microorganisms. 2023 Mar 29;11(4):874. doi: 10.3390/microorganisms11040874. Microorganisms. 2023. PMID: 37110297 Free PMC article.
-
Bacterial Diversity in Old Hydrocarbon Polluted Sediments of Ecuadorian Amazon River Basins.Toxics. 2024 Jan 31;12(2):119. doi: 10.3390/toxics12020119. Toxics. 2024. PMID: 38393214 Free PMC article.
References
-
- Yang Z., Higgs P.I. Myxobacteria: Genomics, Cellular and Molecular Biology. Caister Academic Press; Norfolk, UK: 2014.
-
- Whitworth D.E. Myxobacteria: Multicellularity and Differentiation. ASM Press; Washington, DC, USA: 2008.
-
- Furness E., Whitworth D.E., Zwarycz A. Predatory Interactions between Myxobacteria and Their Prey. In: Jurkevitch E., Mitchell R.J., editors. The Ecology of Predation at the Microscale. Springer; Cham, Switzerland: 2020. pp. 1–36.
Publication types
LinkOut - more resources
Full Text Sources

