Genomic and Chemical Diversity of Bacillus subtilis Secondary Metabolites against Plant Pathogenic Fungi
- PMID: 33622852
- PMCID: PMC8573961
- DOI: 10.1128/mSystems.00770-20
Genomic and Chemical Diversity of Bacillus subtilis Secondary Metabolites against Plant Pathogenic Fungi
Abstract
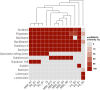
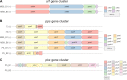
Bacillus subtilis produces a wide range of secondary metabolites providing diverse plant growth-promoting and biocontrol abilities. These secondary metabolites include nonribosomal peptides with strong antimicrobial properties, causing either cell lysis, pore formation in fungal membranes, inhibition of certain enzymes, or bacterial protein synthesis. However, the natural products of B. subtilis are mostly studied either in laboratory strains or in individual isolates, and therefore, a comparative overview of secondary metabolites from various environmental B. subtilis strains is missing. In this study, we isolated 23 B. subtilis strains from 11 sampling sites, compared the fungal inhibition profiles of wild types and their nonribosomal peptide mutants, followed the production of targeted lipopeptides, and determined the complete genomes of 13 soil isolates. We discovered that nonribosomal peptide production varied among B. subtilis strains coisolated from the same soil samples. In vitro antagonism assays revealed that biocontrol properties depend on the targeted plant pathogenic fungus and the tested B. subtilis isolate. While plipastatin alone is sufficient to inhibit Fusarium spp., a combination of plipastatin and surfactin is required to hinder growth of Botrytis cinerea Detailed genomic analysis revealed that altered nonribosomal peptide production profiles in specific isolates are due to missing core genes, nonsense mutation, or potentially altered gene regulation. Our study combines microbiological antagonism assays with chemical nonribosomal peptide detection and biosynthetic gene cluster predictions in diverse B. subtilis soil isolates to provide a broader overview of the secondary metabolite chemodiversity of B. subtilis IMPORTANCE Secondary or specialized metabolites with antimicrobial activities define the biocontrol properties of microorganisms. Members of the Bacillus genus produce a plethora of secondary metabolites, of which nonribosomally produced lipopeptides in particular display strong antifungal activity. To facilitate the prediction of the biocontrol potential of new Bacillus subtilis isolates, we have explored the in vitro antifungal inhibitory profiles of recent B. subtilis isolates, combined with analytical natural product chemistry, mutational analysis, and detailed genome analysis of biosynthetic gene clusters. Such a comparative analysis helped to explain why selected B. subtilis isolates lack the production of certain secondary metabolites.
Keywords: Bacillus subtilis; antiSMASH; biosynthetic gene clusters; chemodiversity; fungal inhibition; secondary metabolites.
Copyright © 2021 Kiesewalter et al.
Figures



Similar articles
-
Phylogenetic Distribution of Secondary Metabolites in the Bacillus subtilis Species Complex.mSystems. 2021 Mar 9;6(2):e00057-21. doi: 10.1128/mSystems.00057-21. mSystems. 2021. PMID: 33688015 Free PMC article.
-
Lipopeptide mediated biocontrol activity of endophytic Bacillus subtilis against fungal phytopathogens.BMC Microbiol. 2019 Apr 2;19(1):71. doi: 10.1186/s12866-019-1440-8. BMC Microbiol. 2019. PMID: 30940070 Free PMC article.
-
Secondary metabolites of Bacillus subtilis impact the assembly of soil-derived semisynthetic bacterial communities.Beilstein J Org Chem. 2020 Dec 4;16:2983-2998. doi: 10.3762/bjoc.16.248. eCollection 2020. Beilstein J Org Chem. 2020. PMID: 33335606 Free PMC article.
-
Biological control of plant pathogens by Bacillus species.J Biotechnol. 2018 Nov 10;285:44-55. doi: 10.1016/j.jbiotec.2018.07.044. Epub 2018 Aug 30. J Biotechnol. 2018. PMID: 30172784 Review.
-
Diversity and Distribution of Volatile Secondary Metabolites Throughout Bacillus subtilis Isolates.Front Microbiol. 2020 Apr 8;11:559. doi: 10.3389/fmicb.2020.00559. eCollection 2020. Front Microbiol. 2020. PMID: 32322244 Free PMC article. Review.
Cited by
-
The Extracellular Lipopeptides and Volatile Organic Compounds of Bacillus subtilis DHA41 Display Broad-Spectrum Antifungal Activity against Soil-Borne Phytopathogenic Fungi.J Fungi (Basel). 2023 Jul 28;9(8):797. doi: 10.3390/jof9080797. J Fungi (Basel). 2023. PMID: 37623568 Free PMC article.
-
Inhibitory activity of bacterial lipopeptides against Fusarium oxysporum f.sp. Strigae.BMC Microbiol. 2024 Jun 27;24(1):227. doi: 10.1186/s12866-024-03386-2. BMC Microbiol. 2024. PMID: 38937715 Free PMC article.
-
Effects of Trichoderma harzianum and Bacillus subtilis on the root and soil microbiomes of the soybean plant INTACTA RR2 PRO™.Front Plant Sci. 2024 Aug 27;15:1403160. doi: 10.3389/fpls.2024.1403160. eCollection 2024. Front Plant Sci. 2024. PMID: 39258296 Free PMC article.
-
Genome sequence of Bacillus pumilus RI06-95 isolated during a Microcystis bloom in lake Champlain, USA.Microbiol Resour Announc. 2024 Oct 10;13(10):e0071324. doi: 10.1128/mra.00713-24. Epub 2024 Aug 30. Microbiol Resour Announc. 2024. PMID: 39212350 Free PMC article.
-
Reduction of Mycotoxigenic Fungi Growth and Their Mycotoxin Production by Bacillus subtilis QST 713.Toxins (Basel). 2022 Nov 17;14(11):797. doi: 10.3390/toxins14110797. Toxins (Basel). 2022. PMID: 36422971 Free PMC article.
References
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

