Comparative analysis of knowledge representation and reasoning requirements across a range of life sciences textbooks
- PMID: 25785183
- PMCID: PMC4362633
- DOI: 10.1186/2041-1480-5-51
Comparative analysis of knowledge representation and reasoning requirements across a range of life sciences textbooks
Abstract
Background: Using knowledge representation for biomedical projects is now commonplace. In previous work, we represented the knowledge found in a college-level biology textbook in a fashion useful for answering questions. We showed that embedding the knowledge representation and question-answering abilities in an electronic textbook helped to engage student interest and improve learning. A natural question that arises from this success, and this paper's primary focus, is whether a similar approach is applicable across a range of life science textbooks. To answer that question, we considered four different textbooks, ranging from a below-introductory college biology text to an advanced, graduate-level neuroscience textbook. For these textbooks, we investigated the following questions: (1) To what extent is knowledge shared between the different textbooks? (2) To what extent can the same upper ontology be used to represent the knowledge found in different textbooks? (3) To what extent can the questions of interest for a range of textbooks be answered by using the same reasoning mechanisms?
Results: Our existing modeling and reasoning methods apply especially well both to a textbook that is comparable in level to the text studied in our previous work (i.e., an introductory-level text) and to a textbook at a lower level, suggesting potential for a high degree of portability. Even for the overlapping knowledge found across the textbooks, the level of detail covered in each textbook was different, which requires that the representations must be customized for each textbook. We also found that for advanced textbooks, representing models and scientific reasoning processes was particularly important.
Conclusions: With some additional work, our representation methodology would be applicable to a range of textbooks. The requirements for knowledge representation are common across textbooks, suggesting that a shared semantic infrastructure for the life sciences is feasible. Because our representation overlaps heavily with those already being used for biomedical ontologies, this work suggests a natural pathway to include such representations as part of the life sciences curriculum at different grade levels.
Keywords: Knowledge representation; Ontology; Question answering; Reasoning; Semantic infrastructure; Textbook knowledge.
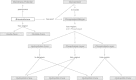
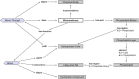
Figures
Similar articles
-
Data-rich textbook figures promote core competencies: Comparison of two textbooks.Biochem Mol Biol Educ. 2021 May;49(3):392-406. doi: 10.1002/bmb.21488. Epub 2021 Jan 9. Biochem Mol Biol Educ. 2021. PMID: 33421340 Free PMC article.
-
Relation-Aware Fine-Grained Reasoning Network for Textbook Question Answering.IEEE Trans Neural Netw Learn Syst. 2023 Jan;34(1):15-27. doi: 10.1109/TNNLS.2021.3089140. Epub 2023 Jan 5. IEEE Trans Neural Netw Learn Syst. 2023. PMID: 34181555
-
Incoming dental students' expectations and acceptance of an electronic textbook program.J Dent Educ. 2011 May;75(5):646-52. J Dent Educ. 2011. PMID: 21546598
-
Thinking Evolutionarily: Evolution Education Across the Life Sciences: Summary of a Convocation.Washington (DC): National Academies Press (US); 2012 Apr 24. Washington (DC): National Academies Press (US); 2012 Apr 24. PMID: 24851293 Free Books & Documents. Review.
-
Mémoire: A framework for semantic interoperability of case-based reasoning systems in biology and medicine.Artif Intell Med. 2006 Feb;36(2):177-92. doi: 10.1016/j.artmed.2005.10.009. Epub 2006 Feb 3. Artif Intell Med. 2006. PMID: 16459063 Review.
References
LinkOut - more resources
Full Text Sources
Other Literature Sources

