CdnL, a member of the large CarD-like family of bacterial proteins, is vital for Myxococcus xanthus and differs functionally from the global transcriptional regulator CarD
- PMID: 20371514
- PMCID: PMC2919716
- DOI: 10.1093/nar/gkq214
CdnL, a member of the large CarD-like family of bacterial proteins, is vital for Myxococcus xanthus and differs functionally from the global transcriptional regulator CarD
Abstract
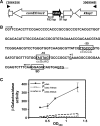
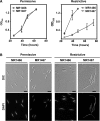
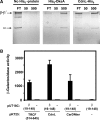
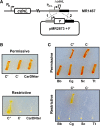
CarD, a global transcriptional regulator in Myxococcus xanthus, interacts with CarG via CarDNter, its N-terminal domain, and with DNA via a eukaryotic HMGA-type C-terminal domain. Genomic analysis reveals a large number of standalone proteins resembling CarDNter. These constitute, together with the RNA polymerase (RNAP) interacting domain, RID, of transcription-repair coupling factors, the CarD_TRCF protein family. We show that one such CarDNter-like protein, M. xanthus CdnL, cannot functionally substitute CarDNter (or vice versa) nor interact with CarG. Unlike CarD, CdnL is vital for growth, and lethality due to its absence is not rescued by homologs from various other bacteria. In mycobacteria, with no endogenous DksA, the function of the CdnL homolog mirrors that of Escherichia coli DksA. Our finding that CdnL, like DksA, is indispensable in M. xanthus implies that they are not functionally redundant. Cells are normal on CdnL overexpression, but divide aberrantly on CdnL depletion. CdnL localizes to the nucleoid, suggesting piggyback recruitment by factors such as RNAP, which we show interacts with CdnL, CarDNter and RID. Our study highlights a complex network of interactions involving these factors and RNAP, and points to a vital role for M. xanthus CdnL in an essential DNA transaction that affects cell division.
Figures




Similar articles
-
Structure-function dissection of Myxococcus xanthus CarD N-terminal domain, a defining member of the CarD_CdnL_TRCF family of RNA polymerase interacting proteins.PLoS One. 2015 Mar 26;10(3):e0121322. doi: 10.1371/journal.pone.0121322. eCollection 2015. PLoS One. 2015. PMID: 25811865 Free PMC article.
-
Structural insights into RNA polymerase recognition and essential function of Myxococcus xanthus CdnL.PLoS One. 2014 Oct 1;9(10):e108946. doi: 10.1371/journal.pone.0108946. eCollection 2014. PLoS One. 2014. PMID: 25272012 Free PMC article.
-
High-mobility-group a-like CarD binds to a DNA site optimized for affinity and position and to RNA polymerase to regulate a light-inducible promoter in Myxococcus xanthus.J Bacteriol. 2013 Jan;195(2):378-88. doi: 10.1128/JB.01766-12. Epub 2012 Nov 9. J Bacteriol. 2013. PMID: 23144251 Free PMC article.
-
The regulatory action of the myxobacterial CarD/CarG complex: a bacterial enhanceosome?FEMS Microbiol Rev. 2010 Sep;34(5):764-78. doi: 10.1111/j.1574-6976.2010.00235.x. Epub 2010 May 20. FEMS Microbiol Rev. 2010. PMID: 20561058 Review.
-
CarD: a new RNA polymerase modulator in mycobacteria.Transcription. 2011 Jan-Feb;2(1):15-8. doi: 10.4161/trns.2.1.13628. Transcription. 2011. PMID: 21326904 Free PMC article. Review.
Cited by
-
Identification of CdnL, a Putative Transcriptional Regulator Involved in Repair and Outgrowth of Heat-Damaged Bacillus cereus Spores.PLoS One. 2016 Feb 5;11(2):e0148670. doi: 10.1371/journal.pone.0148670. eCollection 2016. PLoS One. 2016. PMID: 26849219 Free PMC article.
-
A majority of Rhodobacter sphaeroides promoters lack a crucial RNA polymerase recognition feature, enabling coordinated transcription activation.Proc Natl Acad Sci U S A. 2020 Nov 24;117(47):29658-29668. doi: 10.1073/pnas.2010087117. Epub 2020 Nov 9. Proc Natl Acad Sci U S A. 2020. PMID: 33168725 Free PMC article.
-
Structure-function dissection of Myxococcus xanthus CarD N-terminal domain, a defining member of the CarD_CdnL_TRCF family of RNA polymerase interacting proteins.PLoS One. 2015 Mar 26;10(3):e0121322. doi: 10.1371/journal.pone.0121322. eCollection 2015. PLoS One. 2015. PMID: 25811865 Free PMC article.
-
The conserved transcriptional regulator CdnL is required for metabolic homeostasis and morphogenesis in Caulobacter.PLoS Genet. 2020 Jan 21;16(1):e1008591. doi: 10.1371/journal.pgen.1008591. eCollection 2020 Jan. PLoS Genet. 2020. PMID: 31961855 Free PMC article.
-
CarD integrates three functional modules to promote efficient transcription, antibiotic tolerance, and pathogenesis in mycobacteria.Mol Microbiol. 2014 Aug;93(4):682-97. doi: 10.1111/mmi.12681. Epub 2014 Jul 16. Mol Microbiol. 2014. PMID: 24962732 Free PMC article.
References
-
- Dawid W. Biology and global distribution of myxobacteria in soils. FEMS Microbiol. Rev. 2000;24:403–427. - PubMed
-
- Kaiser D. Signaling in myxobacteria. Annu. Rev. Microbiol. 2004;58:75–98. - PubMed
-
- Zusman DR, Scott AE, Yang Z, Kirby JR. Chemosensory pathways, motility and development in Myxococcus xanthus. Nat. Rev. Microbiol. 2007;5:862–72. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

