Current recommendations for the molecular evaluation of newly diagnosed holoprosencephaly patients
- PMID: 20104604
- PMCID: PMC2815008
- DOI: 10.1002/ajmg.c.30253
Current recommendations for the molecular evaluation of newly diagnosed holoprosencephaly patients
Abstract
Holoprosencephaly (HPE) is the most common structural malformation of the developing forebrain in humans and is typically characterized by different degrees of hemispheric separation that are often accompanied by similarly variable degrees of craniofacial and midline anomalies. HPE is a classic example of a complex genetic trait with "pseudo"-autosomal dominant transmission showing incomplete penetrance and variable expressivity. Clinical suspicion of HPE is typically based upon compatible craniofacial findings, the presence of developmental delay or seizures, or specific endocrinological abnormalities, and is then followed up by confirmation with brain imaging. Once a clinical diagnosis is made, a thorough genetic evaluation is necessary. This usually includes analysis of chromosomes by high-resolution karyotyping, clinical assessment to rule-out well recognized syndromes that are associated with HPE (e.g., Pallister-Hall syndrome, Smith-Lemli-Opitz syndrome and others), and molecular studies of the most common HPE associated genes (e.g., SHH, ZIC2 and SIX3). In this review, we provide current step-by-step recommendations that are medically indicated for the genetic evaluation of patients with newly diagnosed HPE. Moreover, we provide a brief review of several available methods used in molecular diagnostics of HPE and describe the advantages and limitations of both currently available and future tests as they relate to high throughput screening, cost, and the results that they may provide.
2010 Wiley-Liss, Inc.
Figures
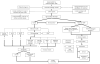
Similar articles
-
New findings for phenotype-genotype correlations in a large European series of holoprosencephaly cases.J Med Genet. 2011 Nov;48(11):752-60. doi: 10.1136/jmedgenet-2011-100339. Epub 2011 Sep 22. J Med Genet. 2011. PMID: 21940735 Free PMC article.
-
Multicolour FISH and quantitative PCR can detect submicroscopic deletions in holoprosencephaly patients with a normal karyotype.J Med Genet. 2006 Jun;43(6):496-500. doi: 10.1136/jmg.2005.037176. Epub 2005 Sep 30. J Med Genet. 2006. PMID: 16199538 Free PMC article.
-
The unfolding clinical spectrum of holoprosencephaly due to mutations in SHH, ZIC2, SIX3 and TGIF genes.Eur J Hum Genet. 2010 Sep;18(9):999-1005. doi: 10.1038/ejhg.2010.70. Epub 2010 Jun 9. Eur J Hum Genet. 2010. PMID: 20531442 Free PMC article.
-
Holoprosencephaly: An update on cytogenetic abnormalities.Am J Med Genet C Semin Med Genet. 2010 Feb 15;154C(1):86-92. doi: 10.1002/ajmg.c.30250. Am J Med Genet C Semin Med Genet. 2010. PMID: 20104602 Review.
-
Holoprosencephaly: from Homer to Hedgehog.Clin Genet. 1998 Mar;53(3):155-63. doi: 10.1111/j.1399-0004.1998.tb02666.x. Clin Genet. 1998. PMID: 9630065 Review.
Cited by
-
A broad range of ophthalmologic anomalies is part of the holoprosencephaly spectrum.Am J Med Genet A. 2011 Nov;155A(11):2713-20. doi: 10.1002/ajmg.a.34261. Epub 2011 Oct 4. Am J Med Genet A. 2011. PMID: 21976454 Free PMC article.
-
Molecular analysis of holoprosencephaly in South America.Genet Mol Biol. 2014 Mar;37(1 Suppl):250-62. doi: 10.1590/s1415-47572014000200011. Genet Mol Biol. 2014. PMID: 24764759 Free PMC article.
-
Middle interhemispheric variant of holoprosencephaly: First prenatal report of a ZIC2 missense mutation.Clin Case Rep. 2020 Apr 30;8(7):1287-1292. doi: 10.1002/ccr3.2896. eCollection 2020 Jul. Clin Case Rep. 2020. PMID: 32695376 Free PMC article.
-
Holoprosencephaly with Clefts: Data of 85 Patients, Treatment and Outcome: Part 1: History, Subdivisions, and Data on 85 Holoprosencephalic Cleft Patients.Ann Maxillofac Surg. 2019 Jan-Jun;9(1):140-145. doi: 10.4103/ams.ams_50_19. Ann Maxillofac Surg. 2019. PMID: 31293943 Free PMC article.
-
TGIF Mutations in Human Holoprosencephaly: Correlation between Genotype and Phenotype.Mol Syndromol. 2010;1(5):211-222. doi: 10.1159/000328203. Epub 2011 May 18. Mol Syndromol. 2010. PMID: 22125506 Free PMC article.
References
-
- Bendavid C, Rochard L, Dubourg C, Seguin J, Gicquel I, Pasquier L, Vigneron J, Laquerriere A, Marcorelles P, Jeanne-Pasquier C, Rouleau C, Jaillard S, Mosser J, Odent S, David V. Array-CGH analysis indicates a high prevalence of genomic rearrangements in holoprosencephaly: an updated map of candidate loci. Hum Mutat. 2009;30:1175–1182. - PubMed
-
- Brown SA, Warburton D, Brown LY, Yu CY, Roeder ER, Stengel-Rutkowski S, Hennekam RC, Muenke M. Holoprosencephaly due to mutations in ZIC2, a homologue of Drosophila odd-paired. Nat Genet. 1998;20:180–183. - PubMed
-
- Bullen PJ, Rankin JM, Robson SC. Investigation of the epidemiology and prenatal diagnosis of holoprosencephaly in the North of England. Am J Obstet Gynecol. 2001;184:1256–1262. - PubMed
-
- Cohen MM., Jr Holoprosencephaly: clinical, anatomic, and molecular dimensions. Birth Defects Res A Clin Mol Teratol. 2006;76(9):658–673. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

